1. Epichaperomics for stressor-to-phenotype investigations:
Development of new strategies to study changes within interactome networks linking stressors to specific phenotypes remains an unmet need. To enable systems level evaluation of proteome-wide changes in PPIs in disease, we developed an ‘omics platform called chemical chaperomics or epichaperomics. Epichaperomics profiling in cancer (Nature 2016), Parkinson’s disease (Nature Comm 2018), and AD (Nature Comm 2020) demonstrated this approach provides insights into human disease biology unavailable through other platforms.
Please click below to watch the VideoByte:
“Epichaperomics can identify disease-related changes in protein-protein interaction networks”

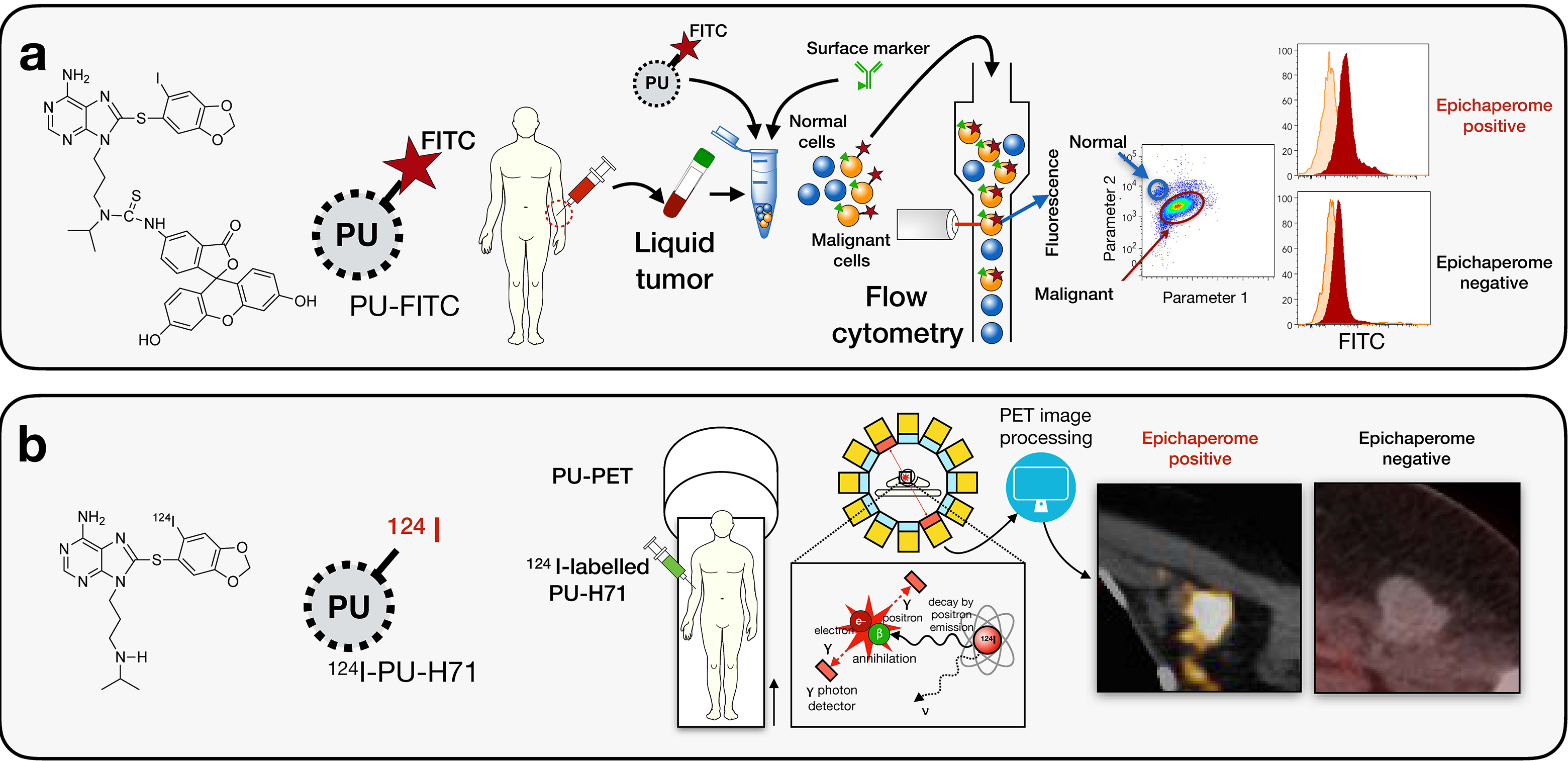
2. Non-invasive assays for detection of the epichaperome in humans.
These assays are based on our finding that PU-H71 and PU-AD specifically interact with the tumor HSP90-incorporating epichaperomes. We have developed a positron emission tomography-based assay that incorporates a radiolabeled version of PU-H71 to detect such epichaperome-positive tumors; and 124I-PU-AD to detect epichaperomes in the AD brain . For liquid tumors we have developed a single-cell flow cytometry-based assay that incorporates a fluorescently labeled PU-H71. The PU-FITC assay for liquid tumors was developed in collaboration with the Guzman lab and the PU-PET assay for solid tumors and the PU-AD PET for Alzheimer’s disease in collaboration with the Lewis and Larson labs. These 3 assays are currently in clinical evaluation.
Rodina A, Xu C, Digwal CS, Joshi S, Patel Y, Santhaseela AR, Bay S, Merugu S, Alam A, Yan P, Yang C, Roychowdhury T, Panchal P, Shrestha L, Kang Y, Sharma S, Almodovar J, Corben A, Alpaugh ML, Modi S, Guzman ML, Fei T, Taldone T, Ginsberg SD, Erdjument-Bromage H, Neubert TA, Manova-Todorova K, Tsou MB, Young JC, Wang T, Chiosis G. Systems-level analyses of protein-protein interaction network dysfunctions via epichaperomics identify cancer-specific mechanisms of stress adaptation. Nat Commun. 2023;14(1):3742. doi: 10.1038/s41467-023-39241-7. https://www.nature.com/articles/s41467-023-39241-7
Sharma S, Kalidindi T, Joshi S, Digwal CS, Panchal P, Burnazi E, Lee SG, Pillarsetty N, Chiosis G. Synthesis of (124)I-labeled epichaperome probes and assessment in visualizing pathologic protein-protein interaction networks in tumor bearing mice. STAR Protoc. 2022;3(2):101318. doi: 10.1016/j.xpro.2022.101318. https://pubmed.ncbi.nlm.nih.gov/35496791/
Alam A, Wang T, Chiosis G. Cytoscape files—Systems-level analyses of protein-protein interaction network dysfunctions via epichaperomics identify cancer-specific mechanisms of stress adaptation [Data set]. Zenodo https://doi.org/10.5281/zenodo.7433980 (2022). https://zenodo.org/record/7433980
Wang T, Digwal CS, Alam A, Chiosis G. R Script Epichaperomics—Systems-level analyses of protein-protein interaction network dysfunctions via epichaperomics identify cancer-specific mechanisms of stress adaptation. Zenodo https://doi.org/10.5281/zenodo.7416220 (2022). https://zenodo.org/record/7416220
