Transmembrane sensing of host immune effectors: The Rip1 and Chalkophore pathways
Signal transduction across membranes through regulated intramembrane proteolysis is conserved throughout all domains of life. In the SREBP pathway of human cells, the membrane-bound SREBP transcription factors undergo sequential cleavage by site-one and site-two proteases (S1P and S2P, respectively), with human S2P being the founding member of the zinc metalloprotease family of intramembrane proteases. RseP, an E. coli S2P, cleaves the anti-Sigma factor for SigE and thereby also regulates transcription in response to environmental signals, specifically unfolded outer membrane proteins. Our original characterization of the Rip1 (Rv2869c) pathway in M. tuberculosis revealed that this S2P is a critical virulence determinant.
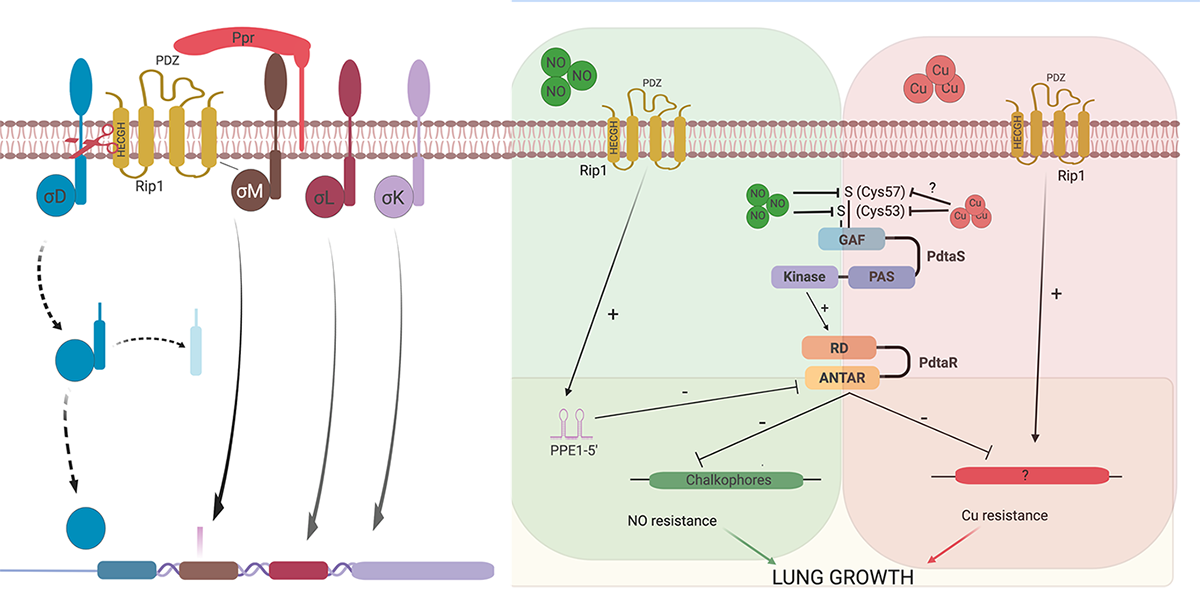
We have identified four anti-Sigma factor substrates of Rip1, anti-SigK, L, M, and D which regulate multiple downstream target genes. However, the virulence functions of Rip1 are independent of these sigma factor pathways. Recent work (Buglino et al, Elife 2021) has identified a new pathway downstream of Rip1 controlled by the PdtaS/PdtaR two component system that controls NO and Cu resistance through a complex set of interacting positive and negative feedback loops. Ongoing work in the lab seeks to understand the molecular details of this signal transduction system that is critical for Mtb virulence.
We have recently focused on the diisonitrile chalkophore system of Mtb. Diisonitrile lipopeptides are produced by Mtb and bind Copper ions with high affinity. Our work has shown that this system allows Mtb to resist copper deprivation both in culture and in the host. This chalkophore acquired copper maintains the copper centers of the Mtb heme:Cu respiratory oxidases, defining a new mechanism by which bacterial pathogens resists host nutritional immunity.
Studies in human TB
In collaboration with our colleagues at Weill Cornell Medicine and the GHESKIO Center in Port Au Prince, Haiti, and with the support of the NIAID’s Tuberculosis Research Unit Program, we are studying the effect of antimycobacterial antibiotics on intestinal microbiome composition and the relationship of those microbiome changes to TB disease resolution.
The Rip1 and Chalkophore pathways
- Makinoshima H. and Glickman M.S., Regulation of Mycobacterium tuberculosis cell envelope composition and virulence by intramembrane proteolysis. Nature, 2005. 436(7049): p. 406-9.
- Sklar, J.G., et al., M. tuberculosis intramembrane protease Rip1 controls transcription through three anti-sigma factor substrates. Mol Microbiol, 2010. 77(3): p. 605-17.
- Schneider JS, Reddy SP, E HY, Evans HW, Glickman MS. Site-2 protease substrate specificity and coupling in trans by a PDZ-substrate adapter protein. Proc Natl Acad Sci U S A. 2013 Nov 26;110(48):19543-8. doi: 10.1073/pnas.1305934110. Epub 2013 Nov 11. PubMed PMID: 24218594; PubMed Central PMCID: PMC3845159.
- Buglino JA, Sankhe GD, Lazar N, Bean JM, Glickman MS. Integrated sensing of host stresses by inhibition of a cytoplasmic two-component system controls M. tuberculosis acute lung infection. Elife. 2021 May 18;10:e65351. doi: 10.7554/eLife.65351. PMID: 34003742; PMCID: PMC8131098.
- Buglino JA, Ozakman Y, Xu Y, Chowdhury F, Tan DS, Glickman MS. 2022. Diisonitrile Lipopeptides Mediate Resistance to Copper Starvation in Pathogenic Mycobacteria. mBio 13:e02513-
- John A Buglino, Yaprak Ozakman, Chad E Hatch, Anna Benjamin, Derek S Tan, Michael S Glickman (2025) Chalkophore-mediated respiratory oxidase flexibility controls M. tuberculosis virulence eLife 14:RP105794. https://doi.org/10.7554/eLife.105794.3
-
Sankhe GD, Xing J, Xiao M, Buglino J, Li H, Zhulin IB, Glickman MS. Ligand binding represses bacterial histidine kinase activity by inhibiting its dimerization. bioRxiv [Preprint]. 2025 Jun 23:2025.05.18.654591. doi: 10.1101/2025.05.18.654591. PMID: 40463091; PMCID: PMC12132213.
The Proteostasis Systems of Mycobacteria
- Fay A, Glickman MS. An essential nonredundant role for mycobacterial DnaK in native protein folding. PLoS Genet. 2014 Jul 24;10(7):e1004516. doi: 10.1371/journal.pgen.1004516. PMID: 25058675; PMCID: PMC4109909.
- Lupoli TJ, Fay A, Adura C, Glickman MS, Nathan CF. Reconstitution of a Mycobacterium tuberculosis proteostasis network highlights essential cofactor interactions with chaperone DnaK. Proc Natl Acad Sci U S A. 2016 Dec 6;113(49):E7947-E7956. doi: 10.1073/pnas.1617644113. Epub 2016 Nov 21. PMID: 27872278; PMCID: PMC5150378.
- Fay A, Czudnochowski N, Rock JM, Johnson JR, Krogan NJ, Rosenberg O, Glickman MS. Two Accessory Proteins Govern MmpL3 Mycolic Acid Transport in Mycobacteria. mBio. 2019 Jun 25;10(3):e00850-19. doi: 10.1128/mBio.00850-19. PMID: 31239378; PMCID: PMC6593404.
- Fay A, Philip J, Saha P, Hendrickson RC, Glickman MS, Burns-Huang K. The DnaK Chaperone System Buffers the Fitness Cost of Antibiotic Resistance Mutations in Mycobacteria. mBio. 2021 Mar 30;12(2):e00123-21. doi: 10.1128/mBio.00123-21. PMID: 33785614; PMCID: PMC8092207.
- Fay A, Kurland AP, Li Z, Monetti M, Johnson JR, Glickman MS.2025.A split ALFA tag-nanobody system for protein localization and proximity proteomics in mycobacteria. mBio16:e0097125.https://doi.org/10.1128/mbio.00971-25
Studies of the role of the microbiome in TB disease resolution with antibiotic therapy
- Wipperman MF, Fitzgerald DW, Juste MAJ, Taur Y, Namasivayam S, Sher A, Bean JM, Bucci V, Glickman MS. Antibiotic treatment for tuberculosis induces a profound dysbiosis of the microbiome that persists long after therapy is completed. Sci Rep. 2017 Sep 7;7(1):10767. doi: 10.1038/s41598-017-10346-6. PMID: 28883399; PMCID: PMC5589918.
- Vorkas CK, Wipperman MF, Li K, Bean J, Bhattarai SK, Adamow M, Wong P, Aubé J, Juste MAJ, Bucci V, Fitzgerald DW, Glickman MS. Mucosal-associated invariant and γδ T cell subsets respond to initial Mycobacterium tuberculosis infection. JCI Insight. 2018 Oct 4;3(19):e121899. doi: 10.1172/jci.insight.121899. PMID: 30282828; PMCID: PMC6237486.
- Wipperman MF, Bhattarai SK, Vorkas CK, Maringati VS, Taur Y, Mathurin L, McAulay K, Vilbrun SC, Francois D, Bean J, Walsh KF, Nathan C, Fitzgerald DW, Glickman MS, Bucci V. Gastrointestinal microbiota composition predicts peripheral inflammatory state during treatment of human tuberculosis. Nat Commun. 2021 Feb 18;12(1):1141. doi: 10.1038/s41467-021-21475-y. PMID: 33602926; PMCID: PMC7892575.
- Bhattarai SK, Du M, Zeamer AL, M Morzfeld B, Kellogg TD, Firat K, Benjamin A, Bean JM, Zimmerman M, Mardi G, Vilbrun SC, Walsh KF, Fitzgerald DW, Glickman MS, Bucci V. Commensal antimicrobial resistance mediates microbiome resilience to antibiotic disruption. Sci Transl Med. 2024 Jan 17;16(730):eadi9711. doi: 10.1126/scitranslmed.adi9711. PMID: 38232140; PMCID: PMC11017772.



