Our work over the past decade has contributed to the detailed genetic and phenotypic characterization of the miR-17~92 and miR-34 families of miRNAs (Ventura et al., Cell 2008; Han, Vidigal, Mu et al., Nature Genetics, 2015, Concepcion et al., PLoS Genetics 2012). We have dissected the oncogenic function of miR-17~92 in lymphomas and prostate cancer (Mu et al., Genes & Dev., 2008) and we have identified, in collaboration with Jeanne Amiel’s group in Paris, hemizygous germline deletions of miR-17~92 as a common genetic cause of Feingold Syndrome, a developmental syndrome characterized by learning disabilities and skeletal defects (De Pontual, Yao, et al., Nature Genetics 2011).
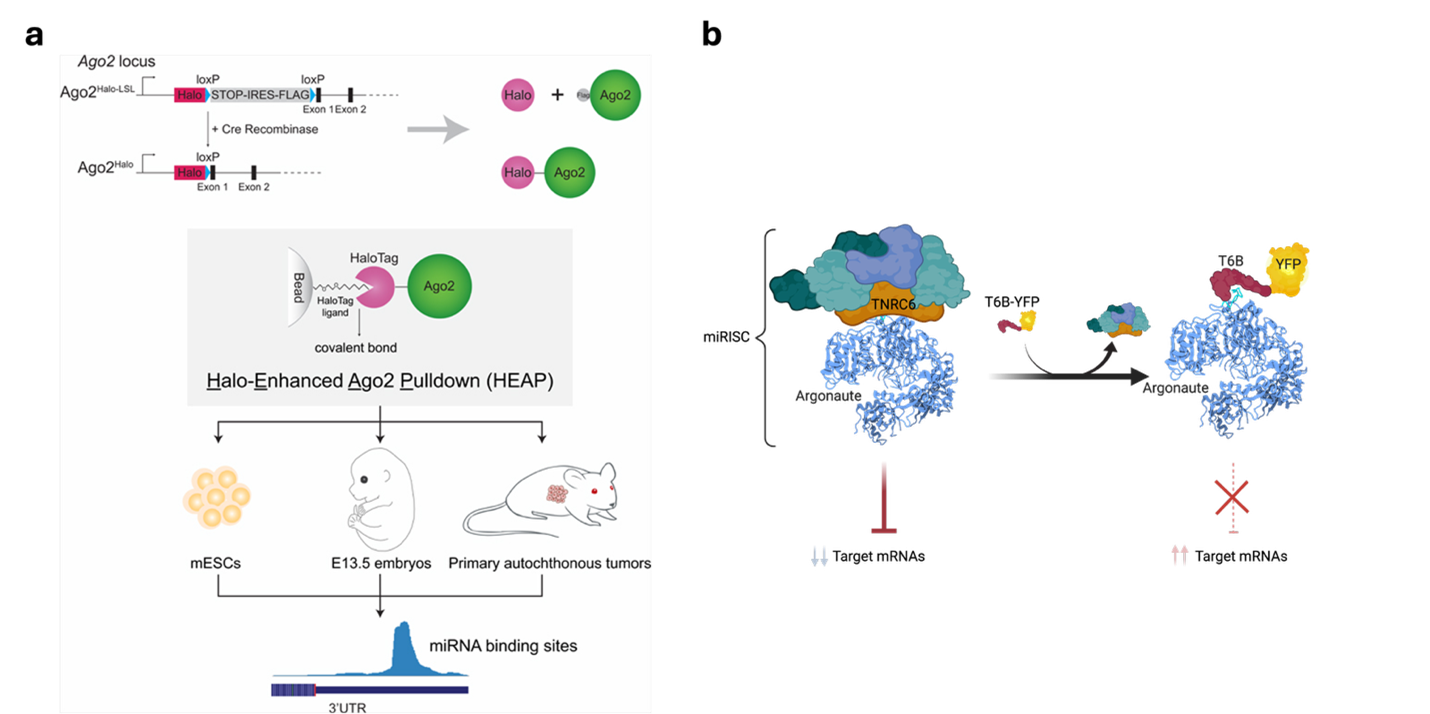
We have also been active in developing better tools to investigate miRNA functions. These include a novel method—Halo Enhanced Ago2 pulldown (HEAP)—based on a genetically engineered mouse strain that allows the direct identification of miRNA/mRNA interaction sites in cells and tissues (Li, Prytikin, Concepcion et al. Mol Cell 2020).
A major challenge in defining the biological functions of miRNAs in adult tissues has been the lack of an effective way to reversibly inhibit their function in a specific, temporally and spatially regulated, fashion. To address this limitation, we have recently generated a novel genetically engineered mouse model harboring a doxycycline-inducible transgene encoding a T6B-YFP fusion protein that, when expressed, prevents the formation of a functional miRISC complex thus acutely blocking miRNA-function. Using this mouse strain, we have shown that although miRNA-mediated gene repression is essential in the skeletal muscle and in the heart, for reasons that we currently do not understand, it is largely dispensable in many other organs and tissues of adult mice under homeostatic conditions. Interestingly, even in these tissues, miRNAs become essential during tissue regeneration and in response to acute tissue damage (La Rocca, King, et al., eLife 2021). More recently, we have explored how miRNA inhibition via T6B can lead to mitotic defects (Zhang, Jiao, et al 2026}.