Two major lines of investigation the lab currently focus on include: modeling human cancers in mice using somatic genome editing and exploring the functional relevance of non-coding RNAs in cancer and development. Below, we detail some ongoing projects:
- How do extrachromosomal circular DNAs (ecDNAs) contribute to tumorigenesis?
- How do chromosomal rearrangements drive tumorigenesis and are these alterations therapeutically actionable?
- How do non-coding RNAs regulate tumorigenesis, regeneration, and homeostasis?
Engineered Extrachromosomal Oncogene Amplifications
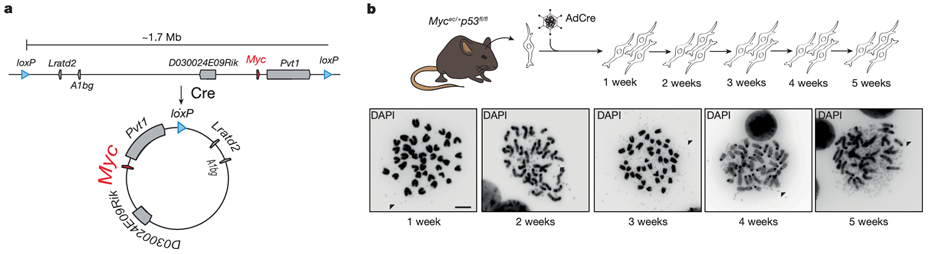
Focal gene amplifications are among the most common cancer-associated mutations but have proven challenging to engineer in primary cells and model organisms. We have recently described a general strategy to engineer large (>1Mb) focal amplifications mediated by extrachromosomal DNAs in a spatiotemporally controlled manner in cells and in mice. We have several mouse models where we can induce ecDNA and we are currently modeling specific tumor types driven by ecDNA, understanding ecDNA evolution, and identifying ways to target ecDNA.
Postdoctoral and pre-doctoral positions are available immediately to join our team and contribute to these efforts—please see Open Positions for more details.
Genetic Screens to Uncover Fundamental Biological Mechanisms
Our lab has developed tools to aid in the generation of guide RNA CRISPR libraries, initially with GuideScan 1.0 (Perez, Pritykin, Vidigal et al., Nature Biotechnology). Since then, we have developed GuideScan 2.0 (Schmidt & Zhang et al., Genome Biology) which is used by labs worldwide (guidescan.com). These tools and others have led us to perform focused and whole-genome CRISPR and shRNA screens.
Leveraging our expertise in genome engineering in both mice and cell lines, we have taken efforts to understand several fundamental molecular mechanisms that drive tumorigenesis including a feedback circuit regulating MYC and the role of co-amplified genes on ecDNAs. Interested members are encouraged to pitch their own screens and areas of investigation.
Postdoctoral and pre-doctoral positions are available immediately to join our team and contribute to these efforts—please see Open Positions for more details.
In vivo and ex vivo chromosome engineering using the CRISPR-Cas9 system
Structural alterations of the genome are commonly observed in human cancer and can contribute to tumorigenesis through a variety of mechanisms including the formation of fusion oncoprotein. The discovery of CRISPR-Cas9 and the development of somatic genome editing methods has profoundly changed the research landscape, providing unprecedented opportunities to faithfully recapitulate the complexity of the cancer genome in mice.
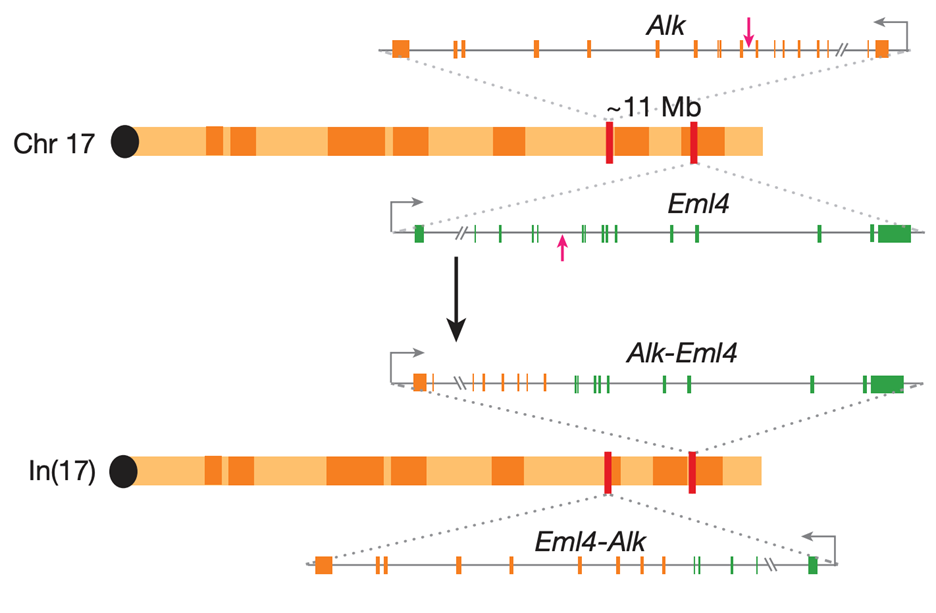
Our group has pioneered the use the CRISPR-Cas9 technology to engineer chromosomal rearrangements in vivo. In 2014, we described a simple strategy to engineer an oncogenic chromosomal inversion leading to the formation of an Eml4-Alk gene fusion in the lungs of adult wild type mice using Sp-Cas9 and 2 gRNAs targeting the desired genomic breakpoints (Maddalo et al., Nature 2014). Since, we have applied this technology to show that a rare intrachromosomal deletion resulting in the formation of BCAN-NTRK1 can drive the formation of highly aggressive gliomas in mice (Cook et al., Nature Communication, 2017). More recently, we have explored two other tumor types driven by chromosomal rearrangements including pilocytic astrocytoma and desmoplastic small round cell tumors.
Roles of non-coding RNAs in Cancer & Development
Over the past two decades, we have contributed to the detailed genetic and phenotypic characterization of the miR-17~92 and miR-34 families of miRNAs in the context of development and cancer (Ventura et al., Cell 2008; Han, Vidigal, Mu et al., Nature Genetics, 2015; Concepcion et al., PLoS Genetics 2012; Mu et al., Genes & Dev., 2008; De Pontual, Yao, et al., Nature Genetics, 2011).
In addition, we have developed better tools to investigate miRNA functions including Halo Enhanced Ago2 pulldown (HEAP) —based on a genetically engineered mouse strain that allows the direct identification of miRNA/mRNA interaction sites in cells and tissues (Li, Prytikin, Concepcion et al. Mol Cell 2020). More recently, we have generated a novel genetically engineered mouse model harboring a doxycycline-inducible transgene encoding a T6B-YFP fusion protein that, when expressed, prevents the formation of a functional miRISC complex thus acutely blocking miRNA-function (La Rocca, King, et al., eLife 2021). More recently, we have explored how miRNA inhibition via T6B can lead to mitotic defects (Zhang, Jiao, et al 2026}.