Structural alterations of the genome are commonly observed in human cancer and can contribute to tumorigenesis through a variety of mechanisms including the formation of fusion oncoprotein and changes in copy number or expression of proto-oncogenes and tumor suppressor genes. Until recently, modeling these structural rearrangements in cells and in whole organisms remained technically challenging, time consuming, and costly.
The discovery of CRISPR-Cas9 and the development of somatic genome editing methods has profoundly changed the research landscape, providing unprecedented opportunities to faithfully recapitulate the complexity of the cancer genome in mice.
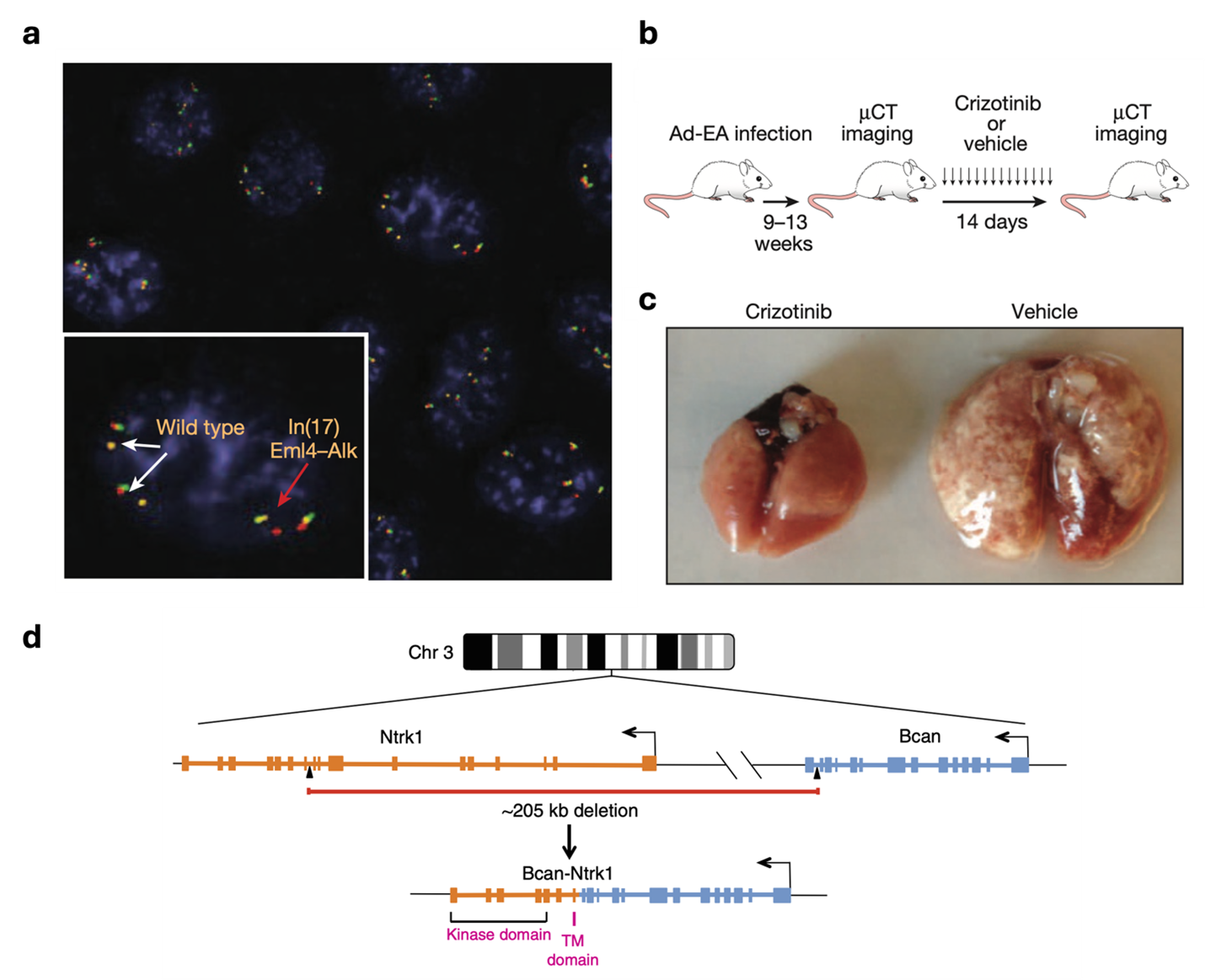
Our group has pioneered the use the CRISPR-Cas9 technology to engineer chromosomal rearrangements in vivo. In 2014 we described a simple strategy to engineer an oncogenic chromosomal inversion leading to the formation of an Eml4-Alk gene fusion in the lungs of adult wild type mice via intra-tracheal delivery of recombinant adenoviruses expressing Sp-Cas9 and 2 gRNAs targeting the desired genomic breakpoints. We showed the this is sufficient to induce the formation of fully penetrant lung adenocarcinoma that are histologically and clinically analogous to human EML4-ALK+ lung cancers (Maddalo et al., Nature 2014).
We have further applied this technology to show that a rare intrachromosomal deletion resulting in the formation of a BCAN-NTRK1 can drive the formation of highly aggressive gliomas in mice (Cook et al., Nature Communication, 2017). Now, we are expanding this approach to study other tumor types including pilocytic astrocytoma (driven by KIAA1549-BRAF) and desmoplastic small round cell tumors (driven by EWRS1-WT1).