Gene regulation by microRNAs
A trove of small RNAs
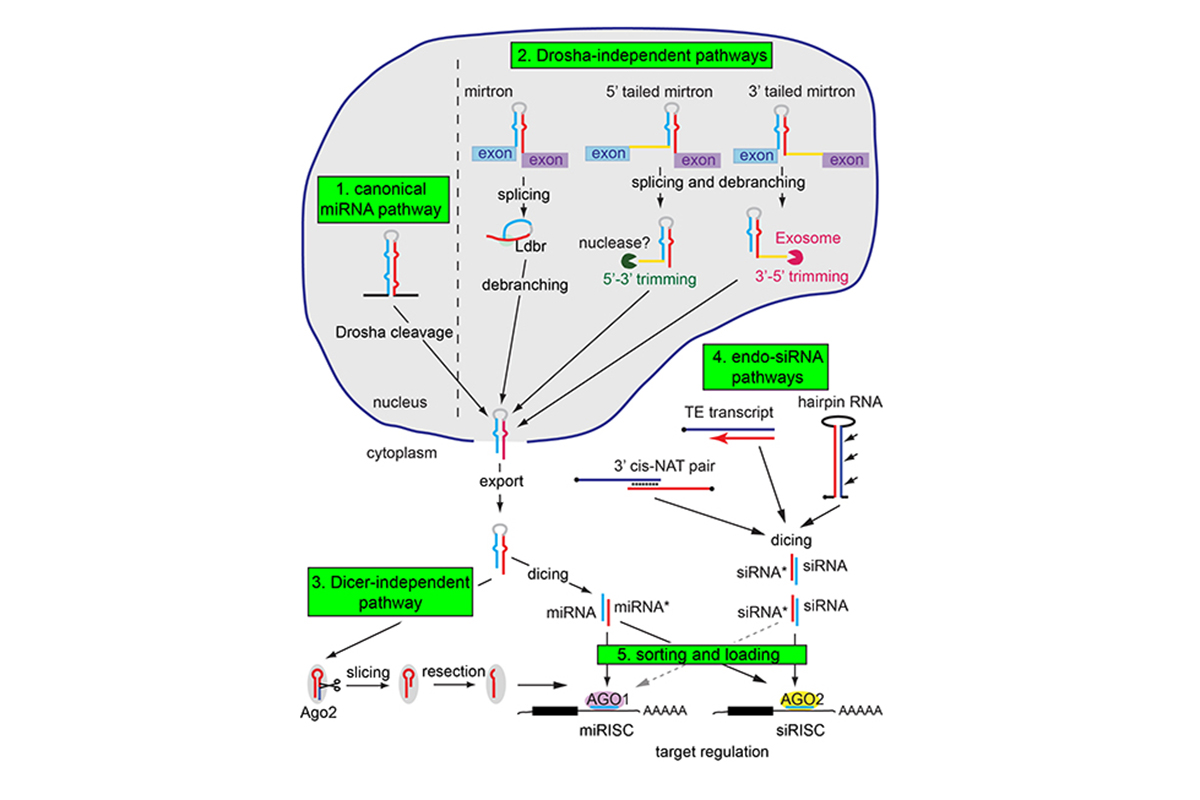
A surprise of the early part of this century was the revelation that small RNAs preside over a previously hidden universe of negative gene regulation. Of particular interest was the discovery that eukaryotic cells process RNAs with double-stranded character into 21-22 nucleotide regulatory RNAs. In the RNA interference (RNAi) pathway, exogenous double-stranded RNA is converted into ~21 nt small interfering (siRNAs), which guide the cleavage and destruction of perfectly complementary transcripts. In a related pathway, endogenous transcripts containing hairpin structures are processed into ~22nt microRNAs (miRNAs) (Figure 1), which negatively regulate host transcripts by cleavage or by translational inhibition. Yet another pathway, termed the piRNA pathway, yields ~24-30 nt RNAs that play critical roles in genome defense, especially in the germline. Understanding the biochemical pathways and mechanistic strategies that produce and mediate the functions of these diverse small RNAs is of great interest, as is elucidating the specific roles of individual small RNA genes, particularly miRNAs and siRNAs.
Biogenesis of diverse small regulatory RNAs
Over the past decade, we have used computational and molecular approaches to annotate mi/si/piRNAs, and to elucidate mechanisms of how they are made and how they are sorted into appropriate Argonaute effector complexes. We have conducted this work in both Drosophila and mammalian systems. Some of the major themes of this work include the following (Figure 1). (1) There turn out to be many “non-canonical” strategies to generate miRNAs, involving unexpected cellular nucleases that normally process other types of transcripts. (2) Loading of different small RNAs into appropriate Argonaute proteins is critical for their appropriate function, and is therefore tightly regulated. (3) There are many steps at which the biogenesis of different small RNA substrates can be regulated by trans-acting factors.
Control of development, behavior and disease by microRNAs
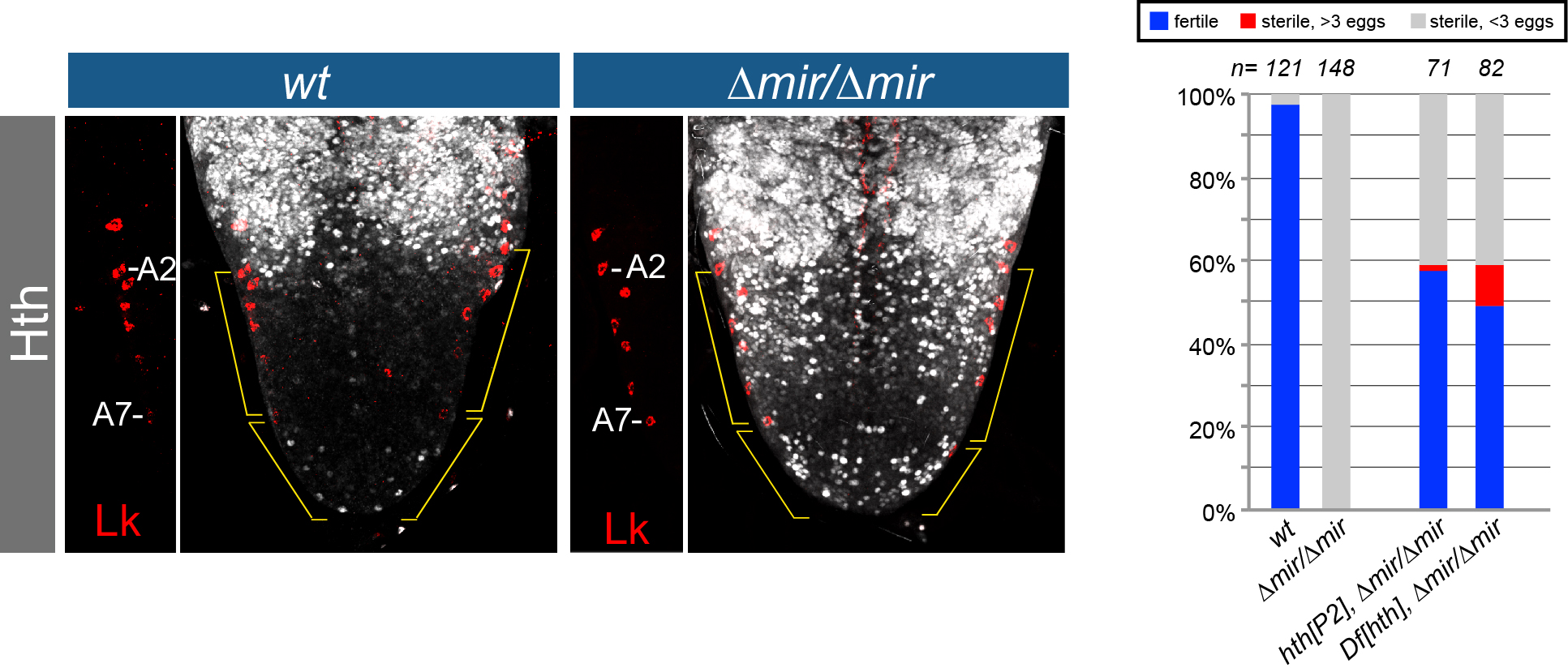
Animal miRNAs generally exhibit only limited complementarity to mRNAs, typically matching to ~7 nt motifs near their 5’ ends (Figure 2). Thus, animal miRNAs tend to regulate large target cohorts. Paradoxically, many animal miRNA knockouts exhibit subtle phenotypes at best, and thus the biological imperative of miRNA regulation can be challenging to infer. This has led to notions that miRNAs are mostly for “fine-tuning” or “robustness”. Nevertheless, the founding miRNAs such as lin-4 and let-7 exhibit profound defects, and our studies have revealed a plethora of developmental and/or behavioral phenotypes in Drosophila miRNA knockouts. Moreover, we have often identified individual mRNA targets whose in vivo deregulation is critical for driving mutant phenotypes. For example, our studies identified transcription factors, RNA binding proteins, and signaling genes as critical miRNA targets during contexts such as development of peripheral sensory organs, wing, eye, and CNS, or during behavioral contexts such as egg-laying, rhythmic behavior, and locomotor activity (Figure 2).
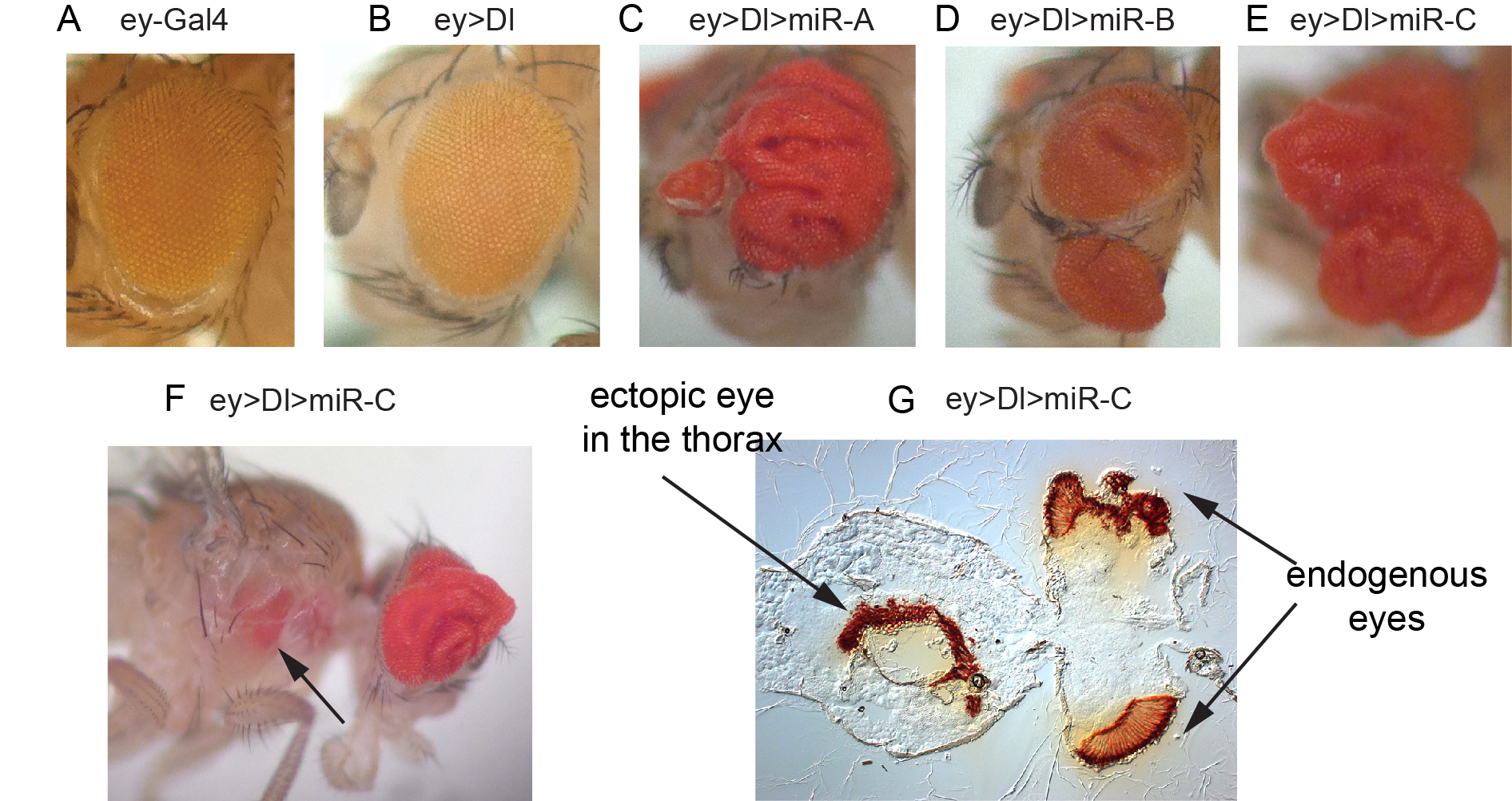
We have also reciprocal in vivo genomewide screens of miRNA activity, which can be uniquely performed in Drosophila using transgenic miRNA libraries that we constructed (Figure 3). We found using unbiased genetic screening that different miRNAs could elicit specific morphological phenotypes that often closely resemble deregulation of known developmental regulators or signaling pathways. In addition, we performed transgenic miRNA modifier screens to uncover miRNAs with unexpected genetic activities in cancer-relevant models. For example, in a screen in the Drosophila eye for modifiers of Notch ligand-dependent growth, we found a variety of miRNAs whose misexpression led to synergistic overgrowth and/or metastatic-like behavior (Figure 3). Dissection of these phenotypes can uncover new principles about in vivo miRNA function and genes that can influence growth and tissue patterning.
Recent work on small RNA biogenesis
Shang R, Kretov D, Adamson S, Treiber T, Treiber N, Vedanayagam H, Chuang JH, Meister G, Cifuentes D and Lai EC (2022). Regulated dicing of pre-mir-144 involves reshaping of its terminal loop. Nucleic Acids Research50: 7637-7654.
Shang R, Baek SC, Kim K, Kim B, Kim VN and Lai EC (2020). Genomic Clustering Facilitates Nuclear Processing of Suboptimal Pri-miRNA Loci. Molecular Cell 16: 303-316.
Vedanayagam J, Chatila WK, Aksoy BA, Majumdar S, Skanderup AJ, Demir E, Schultz N, Sander C and Lai EC (2019). Cancer-associated mutations in DICER1 RNase IIIa and IIIb domains exert similar effects on miRNA biogenesis. Nature Communications 10: 3682.
Jee D, Yang JS, Park SM, Farmer DJ, Wen J, Chou T, Chow A, McManus MT, Kharas MG and Lai EC (2018). Dual strategies for Argonaute2-mediated biogenesis of erythroid miRNAs underlie conserved requirements for Slicing in mammals. Molecular Cell 69: 265-278.
Mohammed J, Flynt AS, Panzarino AM, Mondal M, DeCruz M, Siepel AC and Lai EC (2018). Deep experimental profiling of microRNA diversity, deployment, and evolution across the Drosophila genus. Genome Research 28: 52-65.
Recent work on small RNA biology
Garaulet DL, Moro A and Lai EC (2021). A double-negative gene regulatory circuit underlies the virgin behavioral state. Cell Reports 36, 109335.
Bejarano F, Chang C, Sun K, Hagen JW, Deng WM, and Lai EC (2021). A comprehensive in vivo screen for anti-apoptotic miRNAs indicates broad capacities for oncogenic synergy. Developmental Biology 475: 10-20.
Garaulet DL, Zhang B, Wei L, Li E and Lai EC (2020). miRNAs and Neural Alternative Polyadenylation Specify the Virgin Behavioral State. Developmental Cell 54, 410-423.
Lin CJ, Hu F, Dubruille R, Vedanayagam J, Wen J, Smibert P, Loppin B and Lai EC (2018). The hpRNA/RNAi pathway is essential to resolve intragenomic conflict to permit transmission of sons. Developmental Cell 46: 316-326.
Duan H, de Navas LF, Hu F, Sun K, Mavromatakis YE, Viets K, Zhou C, Kavaler J, Johnston R, Tomlinson A, and Lai EC (2018). The mir-279/996 cluster represses receptor tyrosine kinase signaling to determine cell fates in the Drosophila eye. Development 145: dev159053.
Kavaler J, Duan H, Aradhya R, de Navas LF, Joseph B, Shklyar B and Lai EC (2018). miRNA suppression of a Notch repressor directs non-neuronal fate in Drosophila mechanosensory organs. Journal of Cell Biology 217: 571–583. PMID: 29196461. PMCID: PMC5800810. Featured in JCB Collection on Stem Cells and Development).