In recent years, we decided to adopt our knowledge of studying nuclear DNA stability to the specifics of the mitochondria, and address outstanding questions related to mtDNA replication and repair.
To study mtDNA replication, we developed a novel and powerful single-molecule approach to examine mtDNA replication in vivo, and successfully applied this method to elucidate the mode of mtDNA replication, which has been a matter of controversy for two decades (Phillips et al., Mol Cell 2017). We also defined and for the first time the cause of the common deletion, an mtDNA abnormality that is central to numerous pathologies and aging. Our findings, together with the unique approaches we developed have been very well received by the mitochondrial community. More importantly, they provide a strong foundation for our future work, which aims to address questions related to the physiological impact of mtDNA lesions, including breaks and deletions.
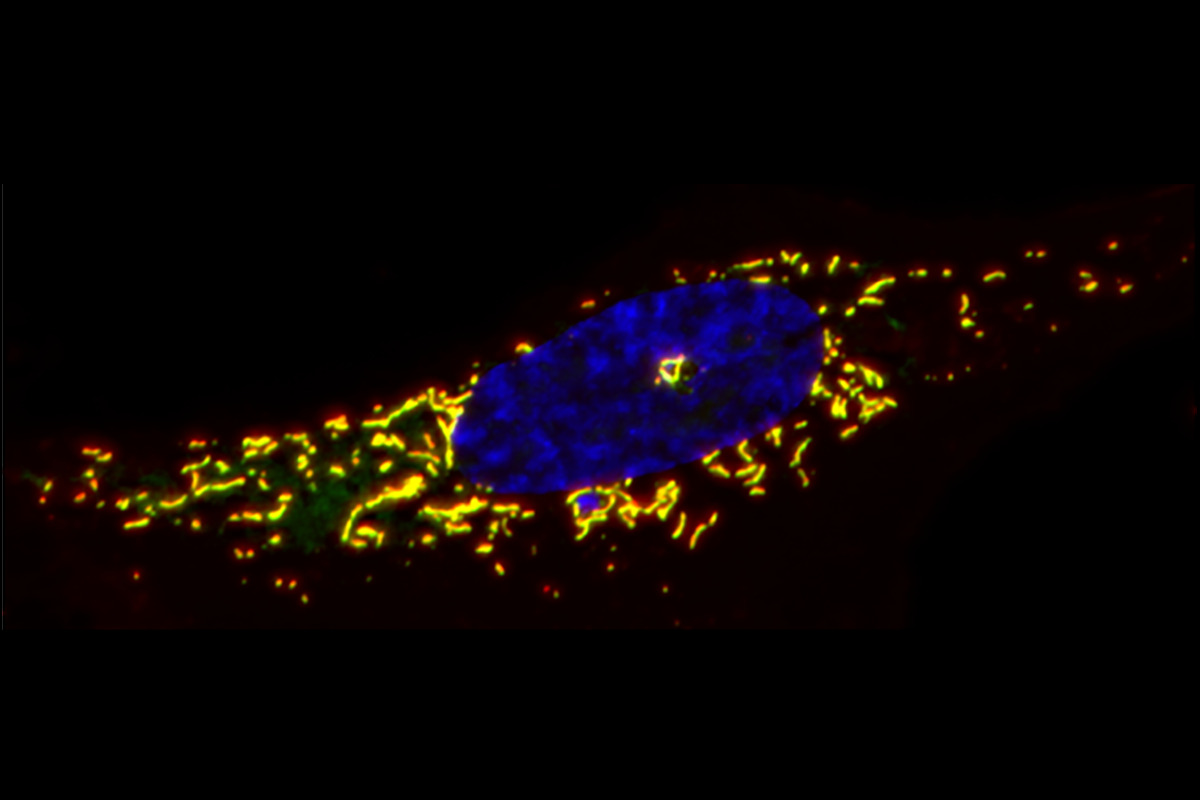
In a recent study, we investigated mitochondrial DSBs (mtDSBs) using mitochondrial TALEN (mTLNs) that we engineered to induce breaks at specific loci within human mtDNA (Tigano et al., Nature 2021). Our work revealed that unlike the nucleus, mitochondria lack robust DSB repair capacity. Instead, broken mitochondrial genomes are predominantly degraded. Subsequent replication of intact mtDNA repopulates mitochondrial genome pool and reestablishes the appropriate copy number. Furthermore, we observed that shortly after mtDSB formation, mammalian cells trigger a type I interferon response as a result of mitochondrial herniation and escape of mitochondrial RNA to the cytoplasm.