As chromosomes go, X and Y make an unlikely pair. The X is large and contains thousands of genes critical for life. The Y, by contrast, is little more than a nub. Its main purpose is to provide the instructions for initiating male development and making sperm. Yet these two very different chromosomes must work together if they are to meet and pair up properly during meiosis — the special form of cell division that creates sperm and egg.
How this happens has remained mysterious for decades. But scientists at the Sloan Kettering Institute have now figured it out. The answer involves some very deliberate breaking and rejoining of DNA.
Breaking is a theme of meiosis. During this process, every chromosome we got from our mothers lines up with every chromosome we got from our fathers and the two swap segments. Before this swapping can occur, the DNA in the chromosomes must be deliberately broken. The regions that are swapped are “homologous” — they’re found in the same place along the chromosome and contain the same genes (although the particular DNA sequence of each gene may be slightly different).
Homologous recombination is vastly more challenging for males because most of the X chromosome has nothing to pair with. In fact, only a very tiny portion of the already tiny Y chromosome has any homology with the X. This region is called the pseudoautosomal region (PAR), and it’s critical for making sure that X and Y find their way into different sperm cells.
Scientists have known for a long time that the PAR undergoes breaking and swapping of segments at a level that far outpaces what one would expect, given its size.
“On most chromosomes, DNA double-strand breaks typically occur once every 10 million base pairs,” says Scott Keeney, a molecular biologist at SKI, who studies this phenomenon. “The PAR in mice is less than 1/10 that size but it still manages to undergo frequent double-strand breaks.”
In a new study published May 27 in the journal Nature, Dr. Keeney and colleagues — including molecular biologist Laurent Acquaviva and longtime collaborator Maria Jasin — show how this happens.
What’s in a Blob?
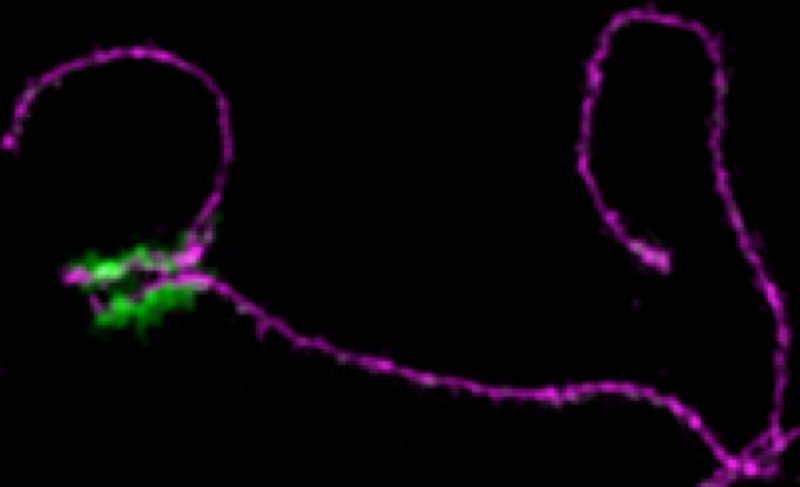
The key to proper pairing of X and Y, they discovered, is a repeated sequence of DNA in the PAR that attracts several double-strand break-related proteins to this region. These protein clusters — which Dr. Acquaviva dubbed “blobs” — change the architecture of the chromosome in this region in such a way that the PAR becomes, as the authors put it, “the hottest area of double-strand break formation in the male mouse genome.”
Similar blobs had been seen in images from published studies. But Dr. Acquaviva — a postdoctoral fellow in the Keeney lab, the lead researcher on the project, and a co-corresponding author on the paper — was the first to define what’s in these blobs and connect them to the hyper-accumulation of double-strand breaks in this region.
“At first glance, the blobs just look like a mess you might see in the microscope if the experiment didn’t work,” Dr. Acquaviva says. “But they turned out to be completely predictable in number, timing, and location, so it became clear that in reality they are very complex structures that the cell builds on purpose.”
In fact, he says, these blobs were key to understanding how the PAR DNA is tethered as short loops to the linear axis that is the structural backbone of the chromosome.
Though the X chromosome also has this same repeated DNA sequence, the two X chromosomes in female meiosis typically do not recombine at this region. Why not? The SKI scientists show that it is because pairing between other regions of the X tends to happen first and directly opposes breakage at the PAR.
This strategy of recruiting more than one’s expected share of DNA-breaking proteins may not be limited to the PAR region. In a paper published earlier this month, the Keeney lab showed that small chromosomes in budding yeast resort to a similar tactic.
Groundbreaking Partnership
These new discoveries, which were made in mice, are the latest fruit of a longstanding collaboration between the Keeney and Jasin labs at SKI. “Scott and I began collaborating back in 1997 when he joined SKI,” she says. “This paper will be our 40th together. It’s a tribute to the collaborative atmosphere of SKI.” In fact, an accompanying editorial published along with the new paper was written by a former collaborative fellow, Francesca Cole, now a faculty member at MD Anderson Cancer Center.
Drs. Jasin and Keeney are both interested in homologous recombination, but they bring complementary expertise to their collaboration. Dr. Jasin is an expert in mammalian double-strand break repair and Dr. Keeney is a specialist in how yeast does meiosis.
Last month, the team’s 39th paper was published in the journal Molecular Cell. In it, they provide the most detailed look yet into how double-strand breaks are repaired. “I am hoping it will change the textbook version of how DNA strands move around during meiotic recombination,” Dr. Jasin says. “But many exciting questions still remain to be tackled.” In other words, stay tuned for more ’breaking’ news from this team of scientists.
This research was supported by the National Institutes of Health (grants F32 GM096692, R35 GM118092, and R35 GM118175), the MSK Cancer Center Support Grant (P30 CA008748), the Lalor Foundation, and the Netherlands Organization for Scientific Research. The study authors declare no competing interests.

