My lab investigates how the genome dictates development using C. elegans as a model.
The Making of a Worm: Every Cell, Every Gene, Every Minute
A central question in modern biology is how the information stored in the genome of a fertilized egg orchestrates the making of an organism. The Nematode C. elegans, aka the worm, is a powerful system to approach the question. The adult worm consists of 959 somatic cells that arise via an invariant cell lineage. The invariant lineage promises single-cell resolution to worm biology, as one can reliably identify any cell at any time by lineage tracing. Exploiting the invariant lineage, our long-term interest is to use the worm as a model to understand how to build a complex organism, gene by gene and cell by cell.
Towards this goal, we have developed a lineaging system that automatically tracks every cell at every minute during worm embryogenesis. The first part of the system is 3D, time-lapse microscopy. We use a ubiquitously expressed histone-GFP fusion to label nuclei and image at one-minute interval over the 14-hour course of embryogenesis.
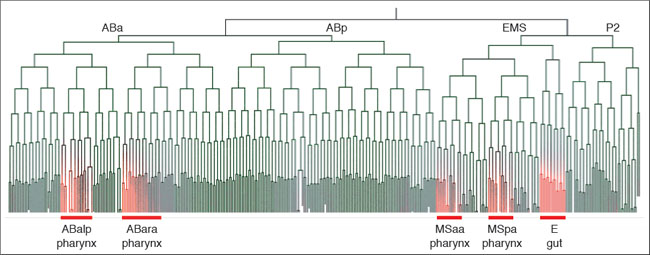
The second part is automated image analysis. We have developed a set of image analysis algorithms to recognize the position and shape of nuclei, track their movements and divisions and construct the lineage (Movie 2). In addition, we have extended the system to 2-color imaging to track gene expression in single cells (Movie 1, Figure 1).
Embryogenesis, or development in general, involves three fundamental processes: proliferation, fate differentiation and morphogenesis. The automated lineaging system enables us to dissect these processes from unique angels of systematic single-cell analysis. In particular, we are asking two questions: (1) how fate controls the pace of proliferation and (2) how cells find their proper migration paths and destination during gastrulation/morphogenesis. In the meantime, we continue to develop new technologies to expand our ability to analyze individual cells as they develop.
Developmental cell cycle control
Through statistical analysis of individual cell cycles in wild type embryogenesis, we found that the pace/length of cell cycle is under tight control by fate throughout the embryo: fate differentiaiton creates small but reproducible differences in division timing among cells that would otherwise divide synchronously. We further found that some known general cell cycle regulators are present in all cells, but only affect the pace of cell cycle in specific lineages. These findings suggest excellent opportunities to study how differentiation pathways interact with the general cell cycle machinery. Because many of these pathways are conserved in other organisms, such studies would shed light on the general principals by which differentiation controls proliferation. In particular, the small but reproducible asynchrony would help us understand how temporal control of development can be so robust yet so sensitive.
A spatial code for morphology?
The invariant lineage of the worm provides a powerful model to study fate patterning, leading to the discovery of various conserved differentiation pathways. Similarly, the invariant body plan and cell positions provide a powerful model to study morphogenesis. We are taking a systematic approach to identify the key cell-cell interactions during gastrulation that specify cell positions and migration paths, using statistical analysis of the wild type embryogenesis, mutant analyses, targeted RNAi screens and laser cell ablation.
Technology Development
We are undertaking two projects to further extend the automated lineaging system. (1) Currently, our image analysis software, StarryNite, can trace through the first 9 out of 10 rounds of embryonic cell division. We are developing new methods to trace through the last round of division, which involves both algorithm development and testing new labels to image cells. (2)We are extending the system to automated, systematic phenotype analysis at single-cell resolution. The lineaging system tracks every cell at every minute in a developing animal with vast information on cell position, migration, division and gene expression. We are developing computational methods to properly extract the information, that is, metrics for automated phenotyping that can serve a wide variety of biological questions. In the long run, we hope to not only track/monitor single cells, but also manipulate them, through lineaging-based automated laser cell ablation, engineering cell-specific genetic switches and other approaches. We would also like to generalize the technologies to other experimental systems, such as Drosophila, mouse and in vitro stem cell differentiation.
We cordially invite enthusiastic scientists to join us and help shape a new lab. Positions are open for a technician/lab manager, a programmer, as well as postdocs and students interested in either or both of the experimental and computational aspects of our research.