The Yueming Li Lab
The main interests of the Li laboratory are to apply integrated approaches of chemistry and biology to address important biological questions and unmet biomedical needs.
Research Interests
Basic & Translational research in Cancer and Alzheimer’s disease:
From disease mechanism to therapeutic development.
Research Areas
Target Identification & Drug Discovery
Chemical Biology & Medicinal Chemistry
Biochemistry & Enzymology
Biophysics & Structural Biology
Animal models & Behavior
Neurobiology & Neuroinflammation
Autophagy, Protein Aggregation & Clearance
γ-secretase
γ-secretase is an intramembrane aspartyl protease complex that has over 150 substrates [1]. γ-secretase is comprised of four subunits: presenilin (the catalytic subunit), nicastrin (responsible for substrate recognition), anterior pharynx-defective 1 (stabilizes the complex), and presenilin enhancer 2 (endoproteolytic activity). These subunits are necessary for the proteolytic activity of γ-secretase [2].
One of γ-secretase’s many cleavage products is Aβ, which ultimately contributes to Alzheimer’s Disease. Amyloid precursor protein (APP) is sequentially cleaved by β- and γ-secretase resulting in a Aβ peptides at various lengths. Aβ42 is considered the most toxic size of Aβ. Mutations in presenilin (the catalytic subunit of γ-secretase) and APP can lead to increases in the Aβ42 to Aβ40 ratio, resulting in Aβ deposition and plaque formation [1, 3].
Another major cleavage product of γ-secretase is Notch intracellular domain (NICD) which translocates into the nucleus to activate target genes [1].
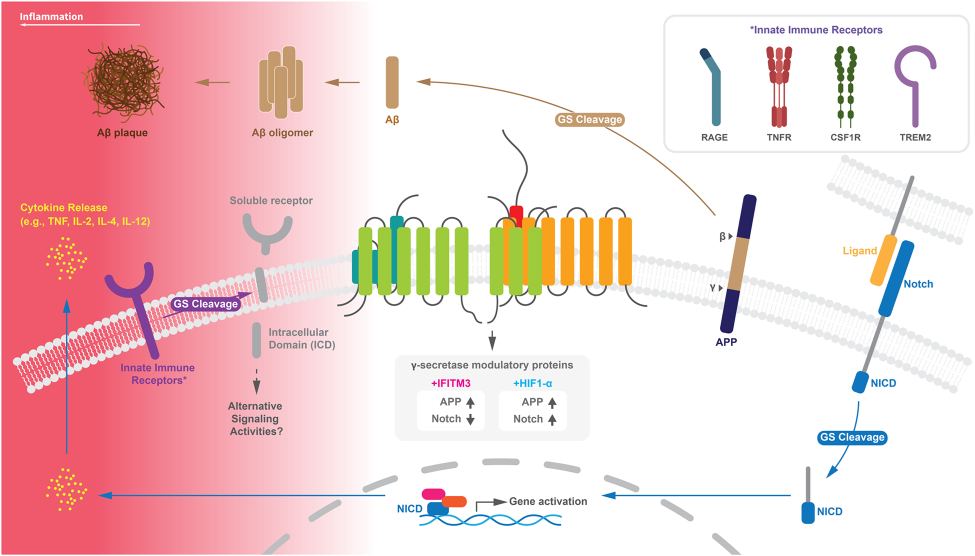
Figure 1: γ-secretase at the center of the innate immunity [4].
Identification of novel γ-secretase modulators
Initially, γ-secretase inhibitors (GSI) were attractive therapeutic targets to prevent the accumulation of Aβ peptides in AD. Due to γ-secretase’s broad substrate specificity, GSIs such as Semagacestat and Avagacestat failed in AD clinical trials [5-7]. Researchers turned to γ-secretase modulators (GSMs): which can alter the cleavage of APP without affecting the rest of γ-secretase’s substrates. GSMs function specifically to prevent the cleavage of APP to Aβ42 without broadly inhibiting γ-secretase’s many other substrates and thereby reducing toxicity [8]. Current work in our lab aims to identify novel GSMs for the treatment of AD.
IFITM3 as a γ-secretase modulatory protein
Our lab identified interferon induced transmembrane protein 3 (IFITM3) as a γ-secretase modulatory protein. In the presence of inflammatory cytokines, IFITM3 directly binds to γ-secretase and upregulates its activity, resulting in increased production of Aβ peptides. The implication of IFITM3 as a γ-secretase modulatory protein marks it as a novel potential therapeutic for AD. IFITM3 knockout resulted in reduced γ-secretase cleavage of APP to Aβ40 and Aβ42 in vitro. This is supported by knockout of IFITM3 in a mouse model of AD (5xFAD), which led to a reduction of both Aβ40, Aβ42, and plaques [9]. An ongoing project aims to identify the cell-specific and temporal roles of IFITM3 by generating conditional knockout cell lines.
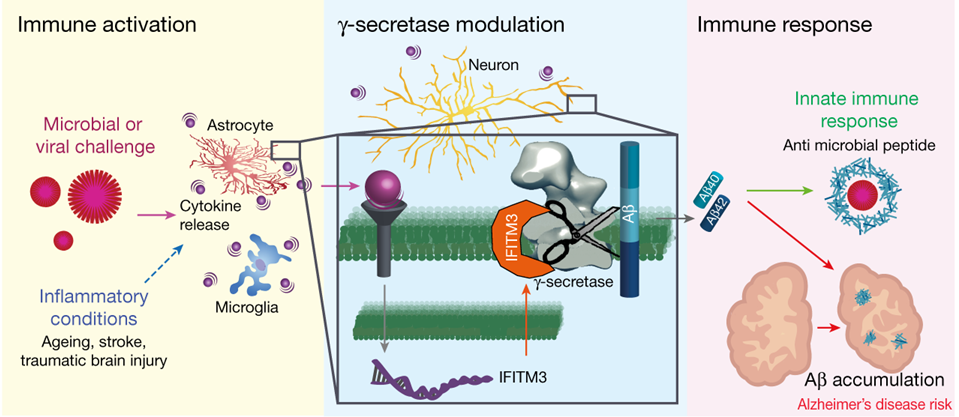
Figure 2: FITM3 connects infections and innate immunity with production of Aβ and Alzheimer’s disease risk [9].
Ferroptosis, a form of cell death triggered by iron-induced lipid peroxidation, is increasingly suspected to contribute to Alzheimer’s disease (AD) progression. However, its precise role in AD pathology remains unclear. As membrane lipid peroxidation is a hallmark of ferroptosis, lipid metabolism and the turnover of membrane phospholipids are being increasingly recognized as key regulators of this process [10]. Astrocytes, as the most abundant brain cells, play critical roles in neuroprotection and brain homeostasis, particularly by regulating lipid metabolism and mitigating oxidative stress [11]. This puts astrocytes as potential key gatekeepers in the brain’s defense or regulation of ferroptosis. It has been shown in-vitro that IFITM3 interacts with cellular cholesterol and modulates membrane lipid composition. Currently, the lab is exploring the interplay between IFITM3, lipid turnover, and ferroptosis in the context of AD pathology.
Tau
Tau aggregation also plays a major role in the development of AD. In healthy neurons, tau stabilizes microtubules. In a diseased state, tau is modified and is prone to aggregation. These insoluble aggregates go on to form neurofibrillary tangles, which are detrimental to neuronal activity and result in cell death. Studying pathogenic tau is key to understanding tau’s role in AD as well as developing therapeutics. Our lab developed two photoaffinity small molecular probes and a novel method for in situ tissue labeling for the study of tau in AD. These two probes offer unique perspectives for studying tau: Tau-2 specifically labels tau across various cell and mouse models while Tau-4 selectively targets the aggregated, high-molecular-weight pathogenic tau in late-stage AD brain tissue [12].
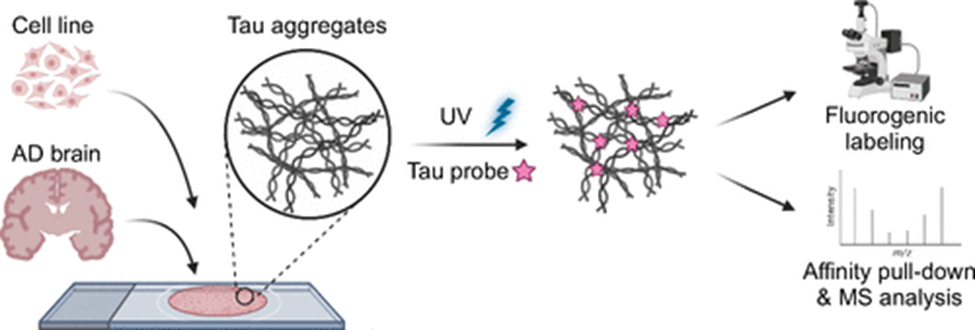
Figure 3: In Situ Labeling of Pathogenic Tau Using Photo-Affinity Chemical Probes [12].
Non-transcriptional Role of HIF-1α
Our lab has uncovered a novel non-transcriptional role of HIF-1α interacting with and activating γ-secretase. This interaction between HIF-1α and γ-secretase has been shown by our lab to increase Notch signaling in breast cancer, exacerbating cancer progression by increasing invasion, migration, metastasis and an increase in cancer stem cells. Additionally, we have shown that Hif1α upregulates BACE1 and γ-secretase activity for Aβ production in hypoxic brain conditions. These results implicate Hif1α as a therapeutic target for AD [13].
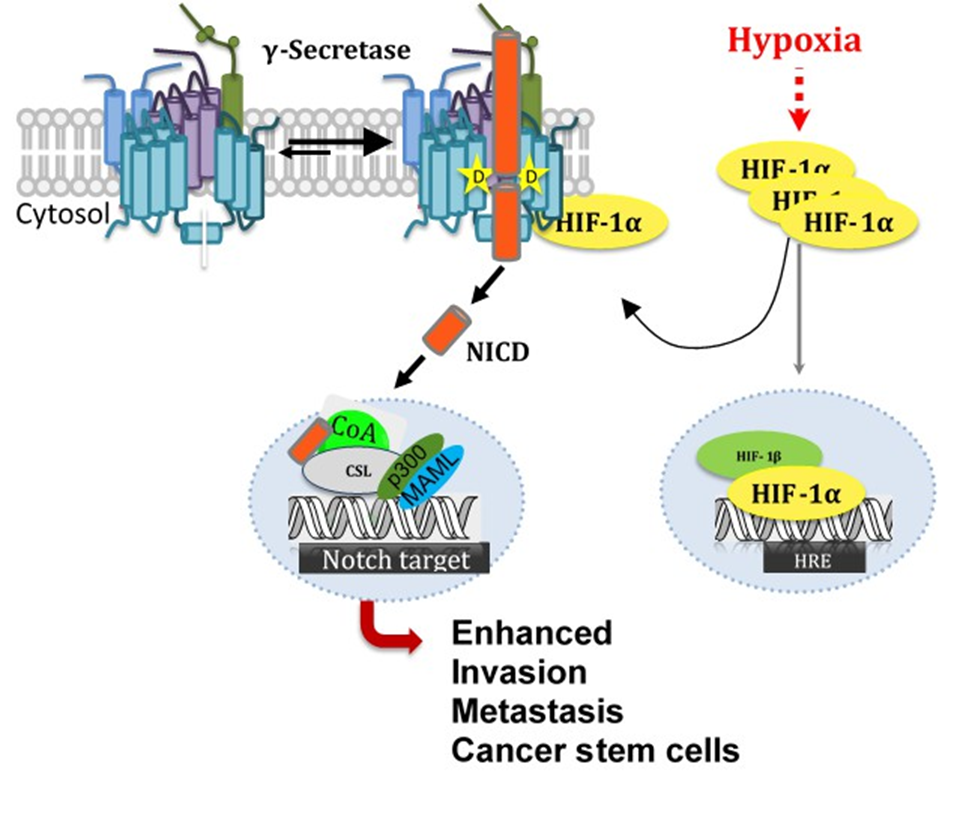
Figure 4: Nontranscriptional Role of Hif-1α in Activation of γ-Secretase and Notch Signaling [14].
Autophagic modulators as a therapy for Alzheimer’s Disease and cancer
Autophagy, a highly regulated pathway, is entered by cells under various conditions including starvation, protein misfolding and oxidative stress. Transcription factor EB (TFEB) has been shown to be the master regulator of the autophagic-lysosomal pathway. Under normal conditions TFEB is phosphorylated by mTORC1 and remains in the cytoplasm. Inactivation of mTORC1 in this pathway thus leads to translocation of TFEB into the nucleus, where it initiates autophagic and lysosomal-related gene expression [15]. Due to increasing reports implicating autophagy in the clearance of tau aggregates in Alzheimer’s disease and in the promotion of tumor progression in various types of cancer, as well as in studies demonstrating the downsides of long-term mTOR, our lab investigates small molecules as autophagic inducers to promote autophagic flux through mTOR independent pathways [16-18]. Our goal is to use autophagic modulators as therapeutic drug treatments for these diseases.
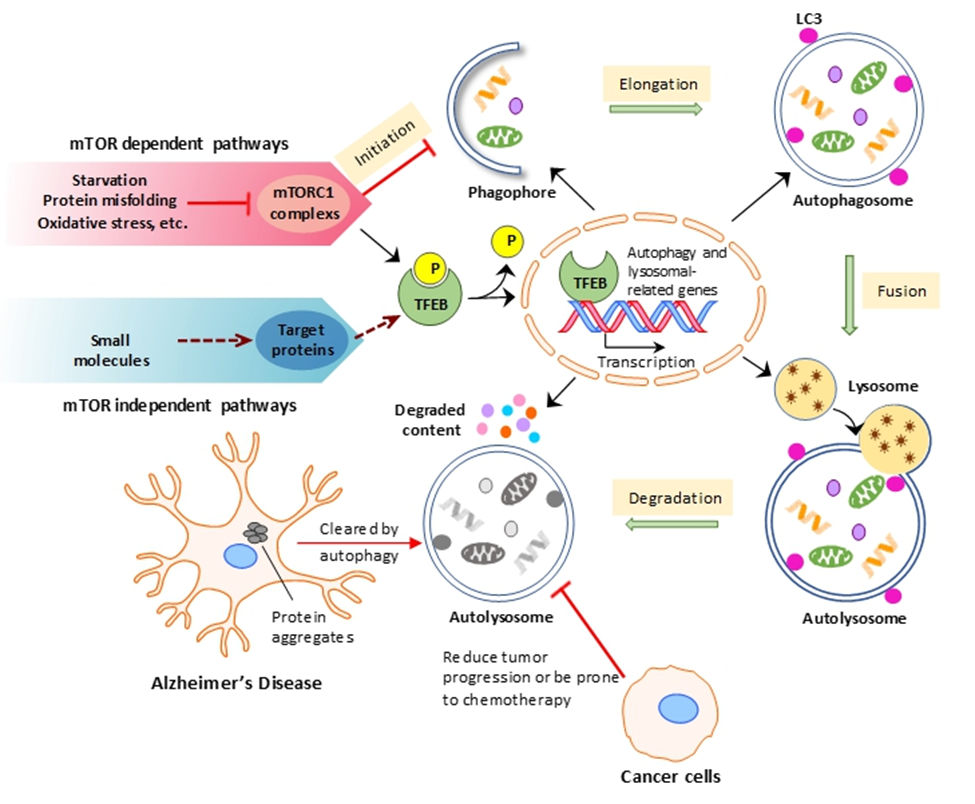
Figure 5: TFEB: the master regulator of the autophagic-lysosomal pathway.
1.Güner, G. and S.F. Lichtenthaler, The substrate repertoire of γ-secretase/presenilin. Seminars in Cell & Developmental Biology, 2020. 105: p. 27-42.
2. Hur, J.Y., gamma-Secretase in Alzheimer’s disease. Exp Mol Med, 2022. 54(4): p. 433-446.
3. Kuperstein, I., et al., Neurotoxicity of Alzheimer’s disease Abeta peptides is induced by small changes in the Abeta42 to Abeta40 ratio. EMBO J, 2010. 29(19): p. 3408-20.
4. Liu, C., C. Nikain, and Y.M. Li, gamma-Secretase fanning the fire of innate immunity. Biochem Soc Trans, 2023. 51(4): p. 1597-1610.
5. Chavez-Gutierrez, L., et al., The mechanism of gamma-Secretase dysfunction in familial Alzheimer disease. EMBO J, 2012. 31(10): p. 2261-74.
6. Crump, C.J., et al., BMS-708,163 targets presenilin and lacks notch-sparing activity. Biochemistry, 2012. 51(37): p. 7209-11.
7. De Strooper, B., Lessons from a failed gamma-secretase Alzheimer trial. Cell, 2014. 159(4): p. 721-6.
8. Crump, C.J., D.S. Johnson, and Y.M. Li, Development and mechanism of gamma-secretase modulators for Alzheimer’s disease. Biochemistry, 2013. 52(19): p. 3197-216.
9. Hur, J.Y., et al., The innate immunity protein IFITM3 modulates gamma-secretase in Alzheimer’s disease. Nature, 2020. 586(7831): p. 735-740.
10. Ma, H., et al., The mechanisms of ferroptosis and its role in alzheimer’s disease. Front Mol Biosci, 2022. 9: p. 965064.
11. Lee, J.A., et al., Lipid metabolism in astrocytic structure and function. Semin Cell Dev Biol, 2021. 112: p. 123-136.
12. Nie, P., et al., In Situ Labeling of Pathogenic Tau Using Photo-Affinity Chemical Probes. ACS Chem Biol, 2025. 20(3): p. 581-591.
13. Alexander, C., et al., Hypoxia Inducible Factor-1alpha binds and activates gamma-secretase for Abeta production under hypoxia and cerebral hypoperfusion. Mol Psychiatry, 2022. 27(10): p. 4264-4273.
14. Villa, J.C., et al., Nontranscriptional role of Hif-1α in activation of γ-secretase and notch signaling in breast cancer. Cell Rep, 2014. 8(4): p. 1077-92.
15. Settembre, C., et al., TFEB links autophagy to lysosomal biogenesis. Science, 2011. 332(6036): p. 1429-33.
16. Jiang, S. and K. Bhaskar, Degradation and Transmission of Tau by Autophagic-Endolysosomal Networks and Potential Therapeutic Targets for Tauopathy. Front Mol Neurosci, 2020. 13: p. 586731.
17. Silva, M.C., et al., Prolonged tau clearance and stress vulnerability rescue by pharmacological activation of autophagy in tauopathy neurons. Nat Commun, 2020. 11(1): p. 3258.
18. Yoon, B.H., et al., Targeted autophagic clearance of Tau protects against Alzheimer’s disease through amelioration of Tau-mediated lysosomal stress. Theranostics, 2025. 15(17): p. 9240-9260.
Lab Links

MetLife Foundation Award Video
The Dance of a Lifetime: Using Creativity and Perseverance in the Art of Science
Is Presenilin-1 really guilty of dismembering Alzheimer protein?
Alz Forum: Probe shines light on gamma-secretase inhibitors
Alz Forum: Yueming Li and Lennart Mucke win MetLife foundation award
Alz Forum: Reconstitution of gamma-secretase