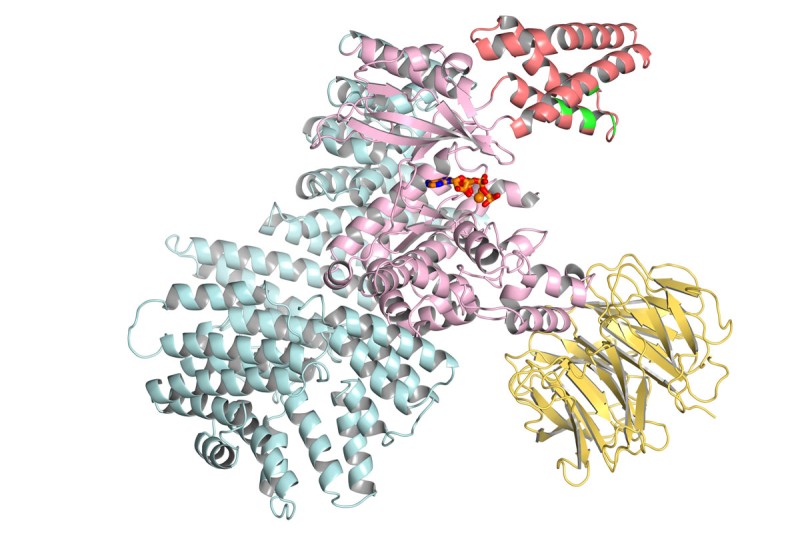
For the first time, structural biologists at Memorial Sloan Kettering have revealed the molecular structure of mTOR, a protein known to be involved in many types of cancer.
The findings, obtained using a method known as x-ray crystallography, provide important insights about how the protein functions. The information may also be used to speed the development of new cancer drugs that act on mTOR.
Three-dimensional renderings of a major portion of mTOR were published in the journal Nature on May 9. In an accompanying commentary, scientists who were not involved in the research observed that the “stunning structure” has been eagerly awaited.
We asked the senior author of the study, Structural Biology Program Chair Nikola P. Pavletich, to tell us more about the findings and their potential value.
What is mTOR, and why are cancer researchers paying close attention to it?
Before a normal cell divides, it senses whether it has enough energy, nutrients, and other factors to grow. The mTOR protein plays a key role in this sensor system by helping the cell receive and respond to a number of external signals.
When growth conditions are favorable, mTOR sets off a cellular process called the mTOR pathway, inducing the cell to grow in size and ultimately causing it to split into two cells.
In many cancers, mutations result in a hyperactive mTOR pathway. As a result, cells may divide excessively and spur the formation of tumors.
How may the structure of mTOR inform cancer research?
Scientists have a fairly good understanding of what signals trigger the mTOR pathway, and how these signals may lead to cancer when the pathway is malfunctioning. In fact, there are hundreds of drugs in development that work by inhibiting the function of mTOR.
But until now, we haven’t fully understood how the protein is regulated at the molecular level – how it is tickled to act.
Once you solve a protein’s structure – in essence, creating a three-dimensional map showing where all its atoms are – you gain much deeper understanding of how the protein works and how its function could be manipulated with drugs.
What challenges did you have to overcome in obtaining the structure?
In x-ray crystallography, you need to produce a protein in large amounts while making it assume its natural shape, and then coax the protein into forming crystals in which all its atoms can be analyzed. In our case, obtaining a crystal wasn’t trivial — partly because mTOR is a huge and complex protein. The only way we could make it fold properly was by producing it in human cells.
Human cells are more difficult to work with than bacteria or insect cells, which are conventionally used for this purpose. A turning point in our research was figuring out how to make human cells produce mTOR in sufficient amounts.
Another challenge was to determine the biochemical conditions needed to work with mTOR in the lab. The way we think of it, it’s learning how to keep the protein happy outside of the cell.
What type of information did the structure reveal?
Like many other cancer-associated proteins, mTOR belongs to a protein type called kinases. We were surprised to learn that the mechanism that controls mTOR activity is different from that of most other kinases and essentially unknown for this type of protein.
By studying the structure of mTOR, we gained a better understanding of how the protein is controlled in normal cells and how it is hyperactivated in cancer.
In addition, we were able to analyze the structure of the mTOR protein bound to drugs that inhibit it. The information we gained could be used to speed the development of these drugs, which hold great promise for cancer treatment.
Could you tell us more about how your findings could help scientists improve upon existing, investigational drugs?
The mTOR-inhibiting drugs that currently are being developed have been discovered though a process of trial and error, which means their specificity and efficacy may not be the best possible.
Knowing what the protein looks like in detail, it is now possible to purposely optimize these drug molecules, a concept known as rational drug design. In the future, we are likely to see new mTOR inhibitors that perform even better than existing drugs — controlling cancer more effectively while minimizing the risk of side effects.
