
This image, generated in the lab of developmental biologist Anna-Katerina (Kat) Hadjantonakis, shows mouse embryos arranged in three circles. The embryos’ appearance varies according to which genes are switched on or off. These developmental genes direct the fate of individual cells during the earliest stage of life.
The image appears on the cover of the June 5 issue of Developmental Cell. Inside, Dr. Hadjantonakis’s team reports an important finding on how genes function during normal mammalian development. Her research, which combines the study of genetics and imaging, is essential for clarifying this process in hopes of putting it to therapeutic use. Guiding undifferentiated cells to become specific cell types could make it possible to replenish those lost to disease or injury.
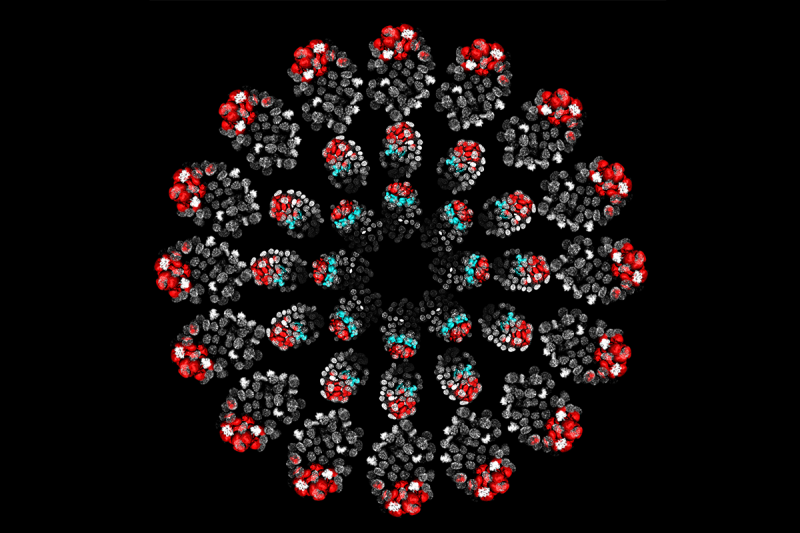
In the middle and outermost rings, the embryo makeup is different — the formation of the blue endoderm layer is either greatly impaired or blocked completely. The researchers engineered these different types of embryos by hindering the effects of either one gene (middle ring) or two genes (outer ring). Both of the blocked genes make receptors that transmit instructions for how a cell should behave.

The receptors are proteins called FGFR1 and FGFR2, and they receive a signal from the FGF4 protein to send embryonic cells down the correct path. (The red and blue coloring is caused by antibodies used to detect the respective proteins.)
Researchers already knew that FGF4 is the main driver of the overall process, but they did not have a clear understanding of its interaction with other proteins.
“We originally thought there was only one receptor receiving the FGF4 signal and that it was expressed only in the cells that become endoderm,” Dr. Hadjantonakis says. “We’ve now shown that the endoderm cells have both FGFR1 and FGFR2, and that they need to act in coordination for the cells to commit to the endoderm state.”
The red pluripotent cells, on the other hand, have just FGFR1. When they receive the FGF4 signal, the cells don’t differentiate — but they undergo a change nevertheless. The cells mature to a more advanced stage of the pluripotent state, a necessary progression before they can ultimately become a specific cell type.
“The FGF4 signal tells one cell type what identity to assume and the other to alter its pluripotent state,” she explains. “This second part was a surprise. We assumed that FGF4 was only directing differentiation, not affecting the pluripotent cells. This finding also reveals that FGFR1 plays a more primary role than FGFR2 in directing cell fate. It’s involved in both transformations.”
Dr. Hadjantonakis is excited about the potential of single-cell imaging and genetic analysis, as well as the recent appointment of Dana Pe’er as Chair of the Computational and Systems Biology Program at the Sloan Kettering Institute.
“She’s one of the pioneers of using computational and mathematical approaches to unravel single-cell data, and she has an expertise complementary to what we are doing in our lab,” Dr. Hadjantonakis says. “I hope that she is as interested and eager for her team to collaborate with us as we are with them.”



