CRISPR-Cas and cGAS-STING Surveillance Pathways
We list below recent highlights on the CRISPR-Cas surveillance pathway.
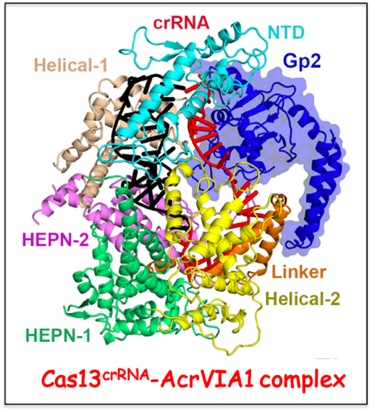
Anti-CRISPR protein targeting type VI CRISPR-Cas complex.
Cas13 recognizes complementary viral transcripts to trigger degradation of both host and viral RNA during the type VI CRISPR-Cas antiviral response. The Luciano Marrafini lab (Rockefeller University) has identified anti-CRISPR protein AcrVIA1 (Gp2) that inactivates the type VIA system from
Listeria seeligeri. We have solved the cryo-EM structure of AcrVIA1 bound to Cas13a
crRNA, establishing that AcrVIA1 interacts with the guide-exposed face of Cas13a, thereby preventing access to the target RNA and the conformational changes required for nuclease activation. Further, unlike other anti-CRISPRs that cause limited immunosuppression, the Marraffini lab showed that a single dose of AcrVIA1 delivered by an individual virion completely dismantles type VI-A CRISPR-mediated immunity.
Meeske, A. J. et al. (2020). Phage-encoded anti-CRISPR enables full escape from type VIA CRISPR-Cas immunity. Science 369, 54-59.
Topology of the type I CRISPR Cascade complexes and targeting by anti-CRISPR proteins.
We have applied cryo-EM in collaboration with the Sriram Subramaniam lab (NCI) to investigate the structure of the type I-F Csy Cascad
crRNA complex in the absence and presence of dsDNA, as well as investigated the impact of anti-CRISPR proteins. Our cryo-EM studies establish ternary and quarternary conformational transitions within Csy Cascade
crRNA on complex formation with dsDNA, events that could contribute to nuclease recruitment and subsequent target degradation.
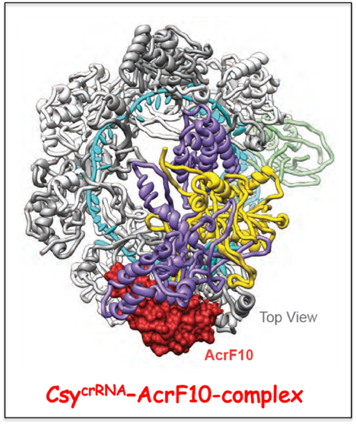
Notably, AcrF10 binds to the PAM DNA recognition site on Cascade
crRNA thereby preventing the first step of target DNA recognition, while AcrF1 binds to two sites on the crRNA of Cascade
crRNA, thereby occluding crRNA-target DNA pairing and R-loop formation.
Guo, T. W. et al., (2017). Cryo-EM structures reveal mechanism and inhibition of DNA targeting by a CRISPR-Cas surveillance complex. Cell 171, 414-426.
Initial step of DNA integration by a CRISPR-Cas transposon complex.
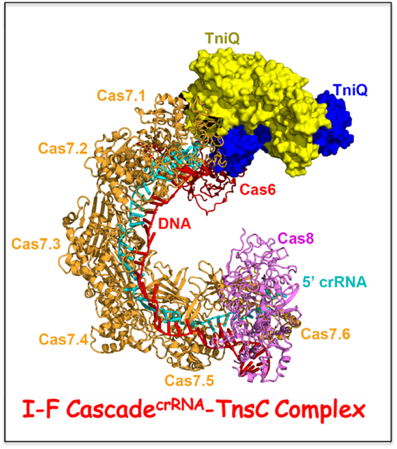
Bioinformatic analysis has revealed the presence of CRISPR-Cas loci in bacterial Tn7-like transposons, thereby implicating a functional relationship between RNA-guided DNA-targeting and transposition, with the latter representing a new role unrelated to host defense. We have solved the cryo-EM structure of the multi-subunit type I-F Cascade
crRNA complex bound to transposition-associated protein TniQ. We find that TniQ binds as a dimer to one end of the Cascade
crRNA complex through contacts with the Cas7.1 and Cas6 subunits, thereby providing insights into the initial step of DNA integration by a CRISPR-Cas-transposition system. Future studies will investigate how subsequent binding of other transposon-associated proteins required for DNA transposition.
Jia, N. et al. (2020). Structure-function insights into the initial step of DNA integration by a CRISPR-Cas-Transposon complex. Cell Research 30, 182-184.
Type III-A CRISPR-Cas System: Regulation of RNA, DNA and cA4-mediated Csm6 Cleavage Activities
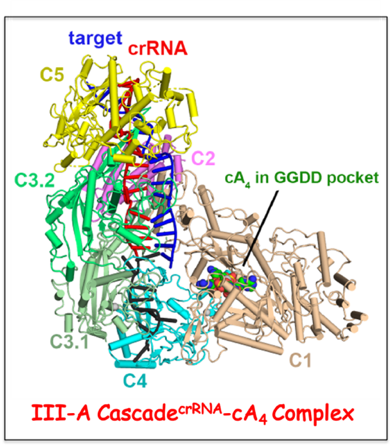
Type III CRISPR-Cas systems provide robust immunity against foreign RNA and DNA by sequence-specific RNase and target RNA-activated sequence-nonspecific DNase and RNase activities. We have solved cryo-EM structures of the type III-A Csm Cascade
crRNA from
Thermococcus onnurineus in the absence and presence of target RNA. The topological features of the crRNA 5
’-repeat tag explains the 5’-ruler mechanism for defining target RNA cleavage sites, with accessibility of positions -2 to -5 within the 5
’-repeat serving as sensors for avoidance of autoimmunity. The Csm3 thumb elements introduce periodic kinks in the crRNA-target RNA duplex, facilitating cleavage of the target RNA with 6-nt periodicity. Key Glu residues within a loop segment positioned over the Csm1 HD pocket of Cascade
crRNA adopt a proposed autoinhibitory conformation suggestive of DNase activity regulation. In addition, we have the formation and positioning of cA
4 in the Csm1 Palm pocket of the Cascade
crRNA, as well as its release path. Finally, we establish cA
4-mediated cleavage of ssRNA by the
trans-acting Csm6 RNase composed of CARF and HEPN domains, utilizing an unanticipated timer mechanism for regulation of the RNase activity of the Csm6 nuclease.
Jia, N. et al. (2019). Type III-A CRISPR Csm complexes: Assembly, target RNA recognition, periodic cleavage and autoimmunity. Mol. Cell 73, 264-267.
Jia, N. et al. (2019). Second messenger cA4 formation within the composite Csm1 Palm pocket of type III-A CRISPR-Cas Csm complex and its release path. Mol. Cell 75, 933-943.
Jia, N. et al. (2019). CRISPR-Cas III-A Csm6 CARF domain is a ring nuclease triggering stepwise cA4 cleavage with ApA>p formation terminating RNase activity. Mol. Cell 75, 944-956.
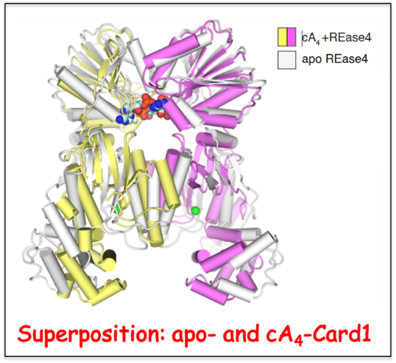
cA4-Mediated activation of Card1 accessory nuclease activity.
The type III CRISPR-Cas immune response produces cyclic oligoadenylate (cA
n) that in turn bind and activate accessory nucleases such as Card1 composed of CARF and REase domains. In a collaborative effort with the Luciano Marraffini lab (Rockefeller University), we have solved the crystal structures of Card1 in the apo- and cA
4-bound states, with the Card1 dimer undergoing a conformational change comprising the rotation of individual subunits relative to each other to form a more compact dimeric scaffold. Activation of Card1 results in the cleavage of both ssRNA and ssDNA
in vitro, with the former cleaved more rapidly than the latter.
In vivo, activation of Card1 induces dormancy of the infected hosts to provide immunity against phage infection and plasmids.
Rostol, J.T. et al. (2021). The Card1 nuclease provides bacterial defense during Type III CRISPR immunity. Nature 590, 624-629.
We list below recent highlights on the cGAS-STING surveillance pathway.
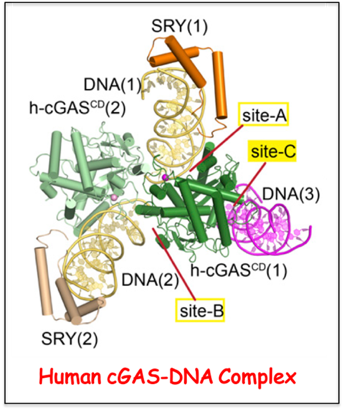
An additional DNA-binding Interface on cGAS contributing to liquid-phase condensation.
The positively charged N-terminal segment of cGAS has been shown by the Zhijian Chen lab (UT Southwestern) to contribute to enhancement of cGAS enzymatic activity as a result of DNA-induced liquid-phase condensation. We have solved the structure of the human cGAS-DNA complex thereby identifying an additional core cGAS
-DNA interface (labeled site-C), such that mutations along this basic interface disrupted liquid-phase condensation. These studies establish that both the N-terminal domain of cGAS and the site-C core cGAS
-DNA interface contribute to DNA-mediated liquid phase condensation. The crystal structure of a triple mutant that disrupts the site C-interface of the human core cGAS
-DNA complex provides a future platform for guiding structure-based cGAS inhibitor development.
Xie, W., Lama, L., Adura, C., Glickman, J. F., Tuschl, T. & Patel, D. J. (2019). Human cGAS catalytic domain has an additional DNA-binding interface that enhances enzymatic activity and liquid phase condensation. Proc. Natl. Acad. Scis. USA 116,11046-11955.
DNA Double Strand Break Repair Pathways
We list below recent highlights on DNA DSB repair pathways.
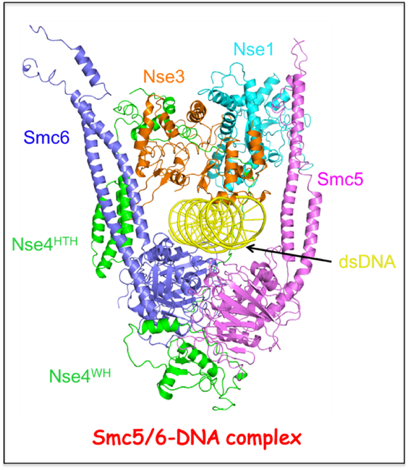
DNA-bound Smc5/6 complex
To address the challenge of how Smc5/6 translocates along DNA, we have solved the cryo-EM structure of DNA-bound 6-subunit
S. cerevisiae Smc5/6 complex, as part of an ongoing collaborative effort with the Xiaolan Zhao lab (MSKCC). A network of interactions between DNA and four of these subunits support the formation of a clamp that encircles the bound DNA double helix. The hook-shaped Nse4 kleisin subunit forms a scaffold connecting DNA and all other subunits. The DNA duplex is positioned above the Smc5 and 6 head regions and between their coiled-coil arm regions, reflecting an ‘engaged-head’ and ‘open-arm’ configuration. An analysis of cross-linking data on apo-Smc5/6 with the structural details for DNA-bound complex has identified multi-subunit conformational changes that enable DNA capture. Future challenges include the structural characterization of complexes containing DNAs lesions and multi-helical junctions, as well as elucidation of the role of Nse5 and6 subunits to the architecture of the complex, the impact of SUMOylation and ubiquitination modifications, and the binding of partner proteins on Smc5/6 function.
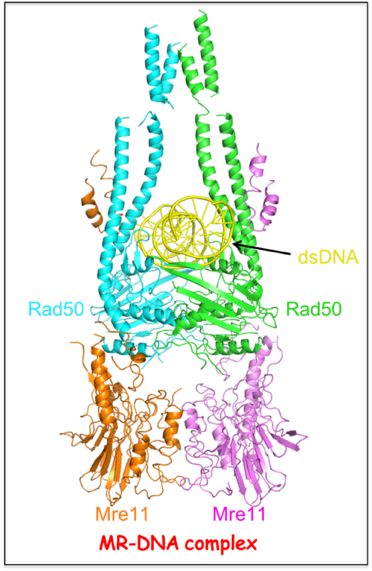
DNA-bound MR complex
DNA double strand breaks (DSBs) can cause severe chromosomal aberrations and genome instability resulting in tumorigenesis. The dimeric MR complex composed of Mre11 nuclease and Rad50 ATPase detects and processes diverse and obstructed DNA ends by initiating DSB repair by either end joining or homologous recombination. To address how the MR complex targets DNA as part of collaborative effort with the John Petrini lab (MSKCC), we have solved the cryo-EM structure of the DNA-bound
S. cerevisiae dimeric MR complex composed of Mre11 and Rad50 subunits. The Rad50 dimer is positioned above the Mre11 dimer with the bound DNA encapsulated within a tunnel through interactions with the head domain and coiled-coil arms of Rad50. This structure has captured an ATP-bound state of MR where the DNA is positioned far from the catalytic nuclease sites of Mre11, and attempts are underway to capture states where the DNA is positioned in proximity to the Mre11 nuclease sites. Additional challenges include structural studies of DNA-bound MRX complex (X stands for the Xrs2 subunit), as well as complexes with MRX’s binding partners towards an understanding its role in DSB repair.
DSB Formation and Repair During Meiosis
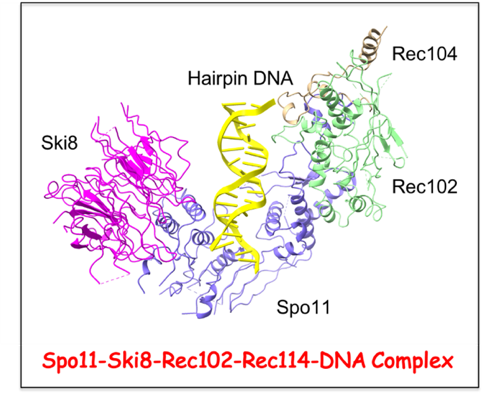
Meiotic recombination initiates with DNA double-strand breaks (DSBs) made by the Spo11 protein together with a suite of accessory factors. In collaboration with the Scott Keeney lab (MSKKC), we are interested in the structural and functional characterization of the DSB Spo11-Rec102–Rec104–Ski8 core complex, as well as the Rec114-Mei4-Mer2 complex that forms nucleoprotein condensates on DNA that are critical for regulating DSB timing, number and location. To this end, we have solved the cryo-EM structure of the
Saccharomyces cerevisiae Spo11-Rec102-Rec104-Ski8 core complex bound to DNA, thereby defining the positioning of the overhang-containing DNA into the Spo11 subunit reflecting formation of the product state following staggered DNA cleavage. Future challenges include structure-function studies of additional states of the DNA-bound core complex, as well as structural elucidation of intermolecular contact patterns stabilizing components of the Rec114-Mei4-Mer2 complex towards a molecular understanding of DSB formation and repair during meiosis.
Readout of Histone and DNA Epigenetic Marks
We list below a recent highlight on histone lysine methyltransferase NSD2 at the nucleosomal level.
Nucleosomal H3K36 methylation by NSD methyltransferase
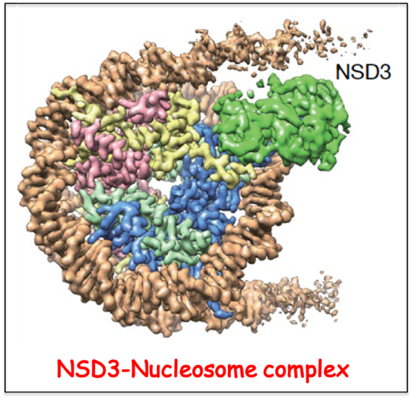
Histone methyltransferases (KMTs) of the nuclear receptor-binding SET domain protein (NSD) family play crucial roles in chromatin regulation and are implicated in oncogenesis. NSD enzymes exhibit an autoinhibitory state that is relieved on binding to nucleosomes, enabling methylation of H3K36. In collaboration with the Zhanxin Wang (Beijing Normal University) lab, we have solved the cryo-EM structures of NSD3 bound to mononucleosomes. We capture unwrapping of the DNA near the linker region, thereby facilitating insertion of the catalytic core between the histone octamer and the unwrapped segment of DNA. A network of DNA- and histone-specific contacts between NSD3 and the nucleosome precisely define the position of the enzyme on the nucleosome, explaining the specificity of methylation to H3K36. In collaboration with the Or Gozani lab (Stanford University), it was shown that several recurrent cancer-associated mutations in NSD3 result from altered intermolecular contacts between NSD proteins and nucleosomes.
Li, W. et al., (2021). Molecular basis of nucleosomal H3K36 methylation by NSD methyltransferases. Nature 590, 498-503.
We list below a recent highlight on maintenance DNA methyltransferase DNMT5.
Cytosine methylation by an ATP-dependent maintenance DNA methyltransferase
Epigenetic evolution occurs over million-year timescales in
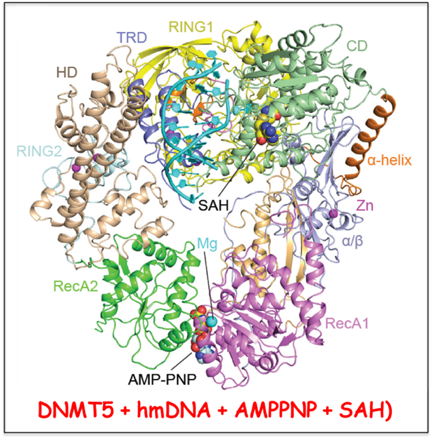
Cryptococcus neoformans and is mediated by DNMT5, a fungal cytosine maintenance methyltransferase dependent on ATP. To understand DNMT5’s novel properties, we solved the cryo-EM structure of
CnDNMT5 in apo-, hemimethylated DNA-bound binary and added ATPgS- and SAH- bound quarternary states. These studies in collaboration with the Hiten Madhani lab (UCal-San Francisco) reveal an elaborate allosteric cascade in which hemimethylated DNA first activates the SNF2 ATPase domain to bind ATP, which then triggers striking structural reconfigurations of both the DNA and the methyltransferase domain, to enable cofactor binding and catalysis, while ejecting non-cognate DNA. This unprecedented chaperone-like, enzyme-remodeling role, of the SNF2 domain within DNMT5 illuminates how energy is used to enable durable nongenetic inheritance outside of the animal and plant kingdoms.
Wang, J. et al. (2022). SNF2 ATPase remodels DNA methyltransferase to enable durable epigenetic memory. Mol. Cell 82, 1186-1198.
Laboratory Personnel and Resources
The Patel laboratory occupies more than 2,000 square feet of contiguous space located on a floor dedicated to structural biology within a modern facility.
Instrumentation Resources
Access to Cryo-EM Resources: The Patel lab has access (time shared with four other groups) to an operational Titan KRIOS with a K3 detector on-site at MSKCC. A second on-site Titan KRIOS with a Falcon4 detector is on order. The Patel lab also has access on a time-share basis (MSKCC is one of nine New York institutions) to four operational 300 kV Titan Krios electron microscopes with K3 detectors at the Simons cryo-EM facility located at the New York Structural Biology Center (NYSBC) in New York city. In addition, the Patel lab has BAG time access to the four Titan KRIOS instruments with K3 detectors at the National cryo-EM Facility located at NYSBC site.
Access to Synchrotron Resources: MSKCC is part of the NE-CAT consortium and the Patel lab has monthly access to their beam lines with micro-focus capabilities at the Advanced Photon Source, Argonne National Laboratory, Chicago, IL. The Patel lab has additional access to operational beam lines at the new NSLS-II synchrotron at the Brookhaven National Laboratory, Long Island, New York.
Access to NMR Resources: The Patel lab has access on a time-share basis to state-of-the-art NMR instrumentation located at NYSBC. This Center has operational Bruker 900 MHz (one each for solution and solids NMR), 800 MHz (three), 700 MHz, 600 MHz and 500 MHz NMR spectrometers each equipped with cryoprobes for solution NMR and one wide bore Bruker 750 MHz spectrometer for solids NMR.
Access to Computational Resources: The Patel lab has two 4-GPU and three 8-GPU servers and 1,400 TB of storage device capacity on-site for processing of cryo-EM data sets. His group also has access to the shared MSKCC Linux cluster with 2,520 CPU cores and 320 NVIDIA GPUs and data storage devices.