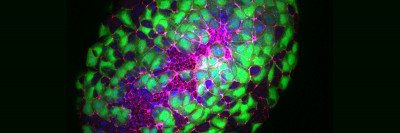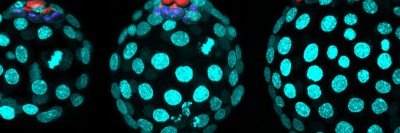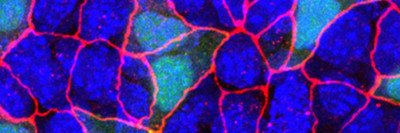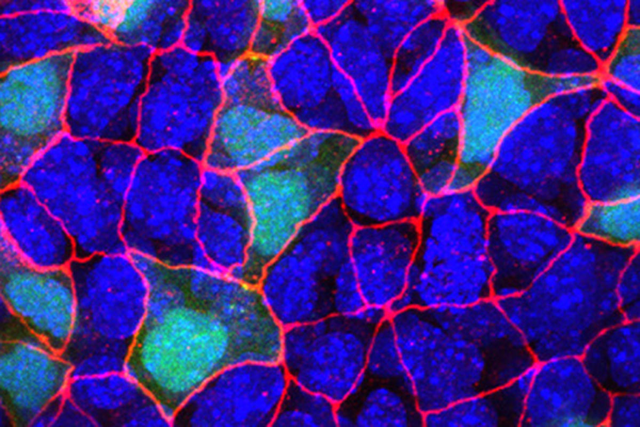
In this fluorescent microscopy image of endoderm tissue from a mouse embryo, cell membranes are red, cell nuclei are blue, and extra-embryonic endoderm cells are green. (They appear turquoise because blue and green are merged.)
To a developmental biologist, animal bodies are a kind of three-layer cake. All of the organs in our bodies, from skin to bones to guts, are made of cells descended from one of three germ layers that form when an embryo is still just a ball of cells.
The outer layer, known as the ectoderm, forms skin, hair, the eyes, and nerves. The middle layer, the mesoderm, forms bones, muscles, and the heart. The innermost layer, the endoderm, makes the majority of our internal organs, including the lungs, liver, pancreas, stomach, intestines, and colon.
Cancers associated with endoderm-derived internal organs are some of the most prevalent, devastating, and hard to treat.
How the cells in an embryo make decisions about their fate — essentially, where they will go and what organs they will become — is a central question of developmental biology. This includes the question of which cells form the body proper and which form the placenta, an extra-embryonic tissue that does not remain with us when we are born.
In a paper published April 8 in the journal Nature, Sloan Kettering Institute investigators present the first-ever complete road map of cells and their fate decisions in an entire germ layer — the mouse endoderm — from fertilization to mid-gestation. The far-reaching analysis, which depended on the joint effort of two research teams, rewrites the story of development.
“The dogma in the field of mammalian development is that the extra-embryonic tissues, such as the placenta, are just that. They’re external to the embryo,” says Kat Hadjantonakis, a developmental biologist in SKI and a co-corresponding author on the paper. “They support the embryo during its development, but they’re dispensable for adult life because the second you are born, you shed your extra-embryonic tissues.”
What she and her co-investigators found, however, is that there is a much more complex trafficking of cells between the extra-embryonic tissues and the embryo. Adult tissues, in fact, contain mixtures of the two.
“The idea that the major internal organs in our body are a mosaic mixture of embryonic and extra-embryonic tissues was shocking to me,” says Dana Pe’er, a computational biologist in SKI and the other corresponding author. “It’s even more surprising than if you told me that the heart was made up of both heart cells and cells from the brain.”
The results raise the question of what these extra-embryonic interlopers are doing and whether they play a specific role in disease. There are also important potential lessons for cancer, since cancers associated with endoderm-derived organs are some of the most prevalent, devastating, and hard to treat.
Mouse Gut Endoderm: Up Close and Personal
The initial observation that the endoderm in a mouse might contain cells from extra-embryonic tissues was made more than a decade ago by Dr. Hadjantonakis’s team. At that time, the discovery elicited more questions than answers.
“One of the questions that always persisted: Are these cells different based on where they come from?” she says. “We could never really address that question because a technique to do so didn’t exist.”
Today, it does. It’s called single-cell RNA sequencing (scRNA-Seq). Using this approach, scientists can isolate single cells from a tissue and then sequence the messenger RNA molecules of it. This provides a readout of all the genes that are turned on in that cell, providing a glimpse into its identity and function.
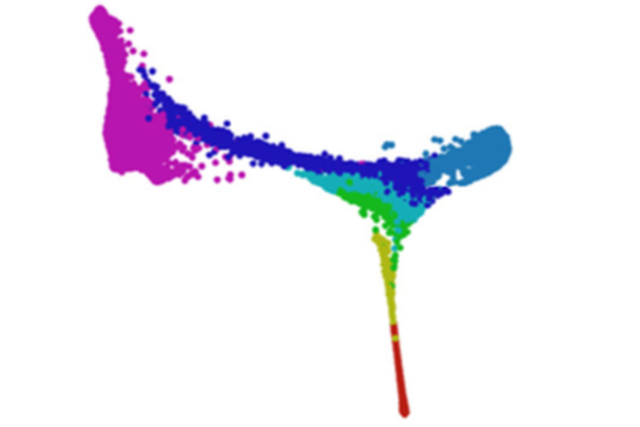
This visual represents the spatial and temporal representation of early development. Each dot represents a single cell and each color represents a different day. The plot demonstrates how cells mature and develop a spatial organization over time in the first days of embryogenesis.
What scRNA-Seq has begun to show in the few years it’s been in use is that there is staggering heterogeneity among cells — even within the same tissue, organ, or tumor — more so than previously realized. All that heterogeneity was lost, or being masked, in approaches that relied on sequencing of pooled mRNA from a larger section of tissue.
Applying this technique — and the computational methods required to interpret the data — to problems of development and cancer attracted Dr. Pe’er to MSK. She arrived in 2016.
Using scRNA-Seq, Dr. Pe’er and her colleagues analyzed gene expression patterns in nearly 120,000 individual cells of the developing mouse embryo. They found that cells that come from extra-embryonic tissues and those that come from the embryo are 99% identical regardless of origin, yet there is a set of genes whose activity is different between them. What’s more, the investigators were able to decipher some of the signals that help cells make their earliest fate decision: whether to become part of the embryo or part of the extra-embryonic structures. The team also pinpointed the earliest time when cells with organ-specific endoderm can be identified. It is far earlier than previously believed.
“Hopefully, we can make use of those differences to follow those cells into later stages of development and adulthood,” says Sonja Nowotschin, a senior research scientist in the Hadjantonakis lab and a co-first author on the paper.
Whether this patchwork of cells with different origins has consequences for the functioning of the organism is an outstanding question. The researchers plan to explore the issue in ongoing work.
Dr. Hadjantonakis hypothesizes that the differences may not matter as much when tissues are healthy, but could have consequences during disease states or periods of regeneration.
A Two-Way Street
While biological questions drove the technical and computational methods used to understand the data, the mathematical tools the group developed ended up pointing to unexpected biological findings. They suggested, for instance, that cells from the embryo actually make their way back out into the extra-embryonic structures and take up residence there.
“At first, the team was skeptical because no one had ever observed cells doing that,” says Manu Setty, a postdoctoral fellow in the Pe’er lab and the other first author on the paper. “But then Sonja designed some experiments in which she looked for the cells in embryos, and sure enough, the computational prediction was right.”
3D reconstruction of the mouse endoderm
Relevance to Cancer
The spirit that motivated the collaboration was a fundamental curiosity about biology, but there are potential implications for medicine.
“One of the things that is being discovered — in my lab and in others — is that cancer cells tend to go back and use these early developmental programs to do their evil deeds,” Dr. Pe’er says.
“Some of the nastiest cancers are those involving internal organs, like the lungs, gut, pancreas, and liver. For combatting them, understanding these early programs in their healthy state is going to be critical,” she adds.