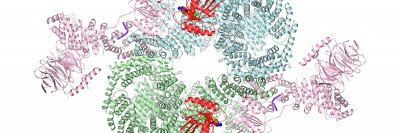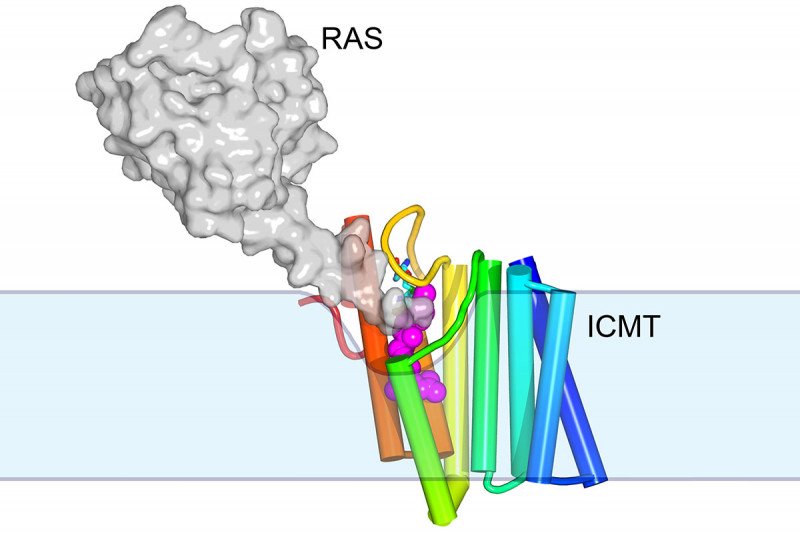
The structure of the membrane protein ICMT, shown in rainbow colors. This protein, an enzyme, attaches a methyl group onto other proteins such RAS (shown in grey) within a membrane (shown in blue).
Every second millions of chemical reactions take place in your cells to kindle the spark of life. Sugars and oxygen combine to produce carbon dioxide, water, and energy. Ions shuttle across membranes, allowing neurons to fire.
Most of these vital chemical reactions would not occur without the help of enzymes. These specialized proteins bring molecules together. The chemical matchmakers help form successful couplings between atoms, speeding the reactions.
Structural biologists know quite a bit about how enzymes cause chemical reactions. But there is one important caveat: Most of what is known concerns soluble enzymes, which float freely in the watery interior of cells. Much less is known about how membrane-embedded enzymes work.
“It’s almost like rewinding the clock 50 years,” says Stephen Long, a structural biologist in the Sloan Kettering Institute who studies these elusive actors. “Even today very little is known about how enzymes catalyze reactions in membranes.”
That’s mostly because membrane proteins are more difficult to crystallize than soluble proteins. Making such crystals is a necessary first step of x-ray crystallography, which is how scientists often determine a protein’s shape. No crystal, no structure.
But newer techniques are finally letting scientists grab hold of membrane proteins and shake loose their mysteries. After ten years of work Dr. Long and his colleagues have determined the structure of an important membrane enzyme called ICMT, which adds methyl groups to proteins. Understanding this structure contributes to our grasp of the biochemistry of membrane enzymes and also suggests ways they could be targeted to treat cancer and other diseases.
Polishing Off Proteins
You can think of ICMT as the last stop on the assembly line for making one specific group of proteins. It adds the final nut — in this case, a methyl group — on the protein, enabling it to function.
The Long lab became interested in ICMT because it’s a good example of a membrane-embedded enzyme that must overcome several obstacles to carry out its function. ICMT itself is hydrophobic, or water fearing. But it must interact with molecules that are hydrophilic, or water loving.
“The situation is not unlike trying to dissolve oil and vinegar in your salad dressing. You can mix them together, but they’ll always separate into different phases,” Dr. Long says.
To accommodate these different binding partners, or substrates, ICMT must have a unique shape that somehow overcomes these hurdles. But determining that shape with x-ray crystallography proved tricky.
Getting Your Bears in Row
To make a good crystal, proteins need to be aligned in a regular and repeating way. Some proteins have a shape that more readily permits such uniformity, while others are less conducive. Dr. Long likens the situation to gummy bears in a box.
“If you shook the box, the gummy bears might all line up with the heads and the arms interlocking in a repetitive pattern,” he says. “But if the bears were more rounded, you can imagine it would be more difficult to pack them all in the same way. You wouldn’t have very consistent interactions between neighboring bears.”
The ICMT enzyme is like a rounded gummy bear. It doesn’t easily form a repetitive arrangement. So Dr. Long’s team first needed to alter its shape slightly. They decided to attach a molecule called a monobody, which is similar to an antibody only much smaller.
“What the monobody does is essentially attach a hat to the enzyme — to the bear,” he says. “The distinctive shape of the hat allows ICMT to make repetitive interactions that form the crystal.”
The other innovative technique the team used is lipidic cubic phase crystallization. With this method, they could crystallize ICMT while it was embedded in a lipidlike environment that mimics the cell membrane.
A former graduate student in the Long lab, Melinda Diver, who is now pursuing a postdoctoral fellowship at the University of California, San Francisco, did all of the crystallization work herself. Dr. Diver tried making crystals out of ICMT proteins from more than 70 species before finding one that worked. The red flower beetle finally did the trick.
Targeting ICMT to Target RAS
The new structure provides tantalizing clues about how membrane enzymes can cause reactions, using substrates located both within and outside of the membrane. “There’s a little groove in the enzyme into which the hydrophobic substrate can diffuse from the membrane,” Dr. Long explains. “And then there’s a tunnel with a little cap that flips open to allow the water-soluble substrate to come in.” The end result is a protein with a methyl group attached.
One of the 200 or so proteins that ICMT modifies in this way is RAS, which is commonly mutated in cancer. Mutated RAS is essentially stuck in the on position. It signals cells to grow without stopping. Targeting ICMT with a drug could be one way to treat RAS-driven cancers. RAS without its methyl group would be less able to plug itself into the cell’s membrane to function.
The team plans to test this hypothesis next by seeing if monobody inhibitors of ICMT can control the growth of RAS-driven tumors in the lab.
There’s also still work to be done to understand exactly how catalysis works at the enzyme’s active site.
“From both therapeutic and biochemical standpoints, we have a lot to do,” Dr. Long says.
A paper describing the new structure is published in the journal Nature.